-
细胞器高度的动态过程和相互作用对复杂细胞功能具有调控作用[1]。真核细胞中的细胞器主要包括细胞核、内质网、高尔基体、溶酶体、线粒体、叶绿体、过氧化物酶体、内吞体、脂滴等,它们以膜为屏障维持特定形态及特定功能,称为膜性细胞器[2]。此外,还有非膜性细胞器,主要包括核糖体和中心体。由于细胞器具有独立的局部环境和独特的分子组成,它们可以独立发挥特定功能,同时各细胞器间相互交流、相互作用,从而协调各自功能的正常发挥,有助于细胞正常行使一系列生理功能[3]。因此,研究细胞器间相互作用对生理病理条件下细胞功能研究具有重要的意义。
细胞器的功能与其组成、结构密切相关,研究生理病理条件下细胞器结构、形态、组成变化规律是研究其功能以及细胞器之间相互作用的基础。研究细胞器的超精细结构和功能,深入探索细胞器间相互作用规律成为生理病理条件下细胞功能研究的关键问题。目前细胞器结构、形态、组成变化及相互作用研究中,最直接的手段为显微成像技术,主要有透射电子显微成像技术、原子力显微成像技术与荧光显微成像技术三种:(1)透射电子显微镜成像具备纳米级空间分辨率,可实现细胞器精细结构、分布及蛋白质等生物大分子成像与分析。然而,电子显微镜无法用于活细胞样品成像,同时电子束在成像过程中对样品形态有一定的影响,从而限制了其在活细胞内细胞器真实结构及相互作用规律研究中的应用;(2)原子力显微成像,分辨率可以达到纳米尺度,样品损伤小,不破坏生物样品的天然形貌,同时能够在液体环境中进行成像,因此它能够用于活细胞结构成像。然而,由于其是通过探针和样品表面的相互作用来进行成像的,只能获得细胞的表面信息,对活细胞内部细胞器结构的研究需要搭配特定的方法才能实现,从而限制了其在细胞器相互作用研究中的应用[4];(3)荧光显微成像是典型的活细胞成像方式。其中,激光扫描共聚焦显微镜空间分辨率较高,可通过细胞器或目标生物分子特异标记荧光探针,实现目标细胞器或分子的可视化成像,其已经成为研究细胞器及其标志分子定位、相互作用的重要工具[5]。然而,激光共聚焦显微镜分辨率受到衍射极限(>200 nm)的限制[6],在深入的研究中已无法满足细胞器结构及标志分子的高精度成像要求,限制了其在细胞器相互作用研究中的进一步应用。
随着成像技术的发展,研究人员开发了超分辨显微成像(Super-resolution microscopy, SRM)技术,突破了衍射极限[7],可以更加清晰的呈现细胞器的精细结构和相互作用位点,成为活细胞研究的重要手段[8]。超分辨显微成像技术可以实现多个细胞器的结构和动力学(运动、相互作用、重塑等)的研究,从而揭示细胞器之间复杂的相互作用,例如膜接触位点(指细胞器相互作用界面,Membrane contact sites, MCS)的形成以及细胞器在细胞内分布的协调[9]。细胞器之间通过MCS蛋白在功能上相关并在结构上锚定,锚定之后的相互作用不仅调节细胞器形态[10],蛋白质修饰、转移、分泌,细胞器动力学[11-12]、脂质合成[13]和钙转移[14]、线粒体分裂与DNA合成,还可以协调不同的细胞器以响应各种细胞刺激,包括改变它们的定位[15]。超分辨显微成像技术逐渐在细胞器精细结构、动态相互作用及形态变化规律研究中发挥重要作用,从而为生理病理现象背后的机制探索提供了新思路与新手段。
基于超分辨显微成像技术在细胞器相互作用研究中应用的快速发展,文中介绍了受激发射损耗(Stimulated emission depletion, STED)显微成像、结构光照明显微成像(Structured illumination microscopy, SIM)、单分子定位显微成像(Single molecule localization microscopy, SMLM)技术,并总结了这几种超分辨显微成像技术在细胞器相互作用中的应用与现状,最后对超分辨显微技术的局限性和发展方向进行总结与展望。
-
超分辨显微成像技术按照其成像特点可分为三类:一是基于压缩点扩散函数的STED显微成像技术 [16];二是SIM技术[17];三是基于荧光分子开关效应的SMLM技术[18],以光激活定位显微成像(Photo activation localization microscopy, PALM)和随机光学重建显微成像(Stochastic optical reconstruction microscopy, STORM)技术为代表。根据庄小威等[19]的综述总结以上超分辨显微成像技术的特点,见表1。
表 1 三类超分辨显微成像技术的原理及特点
Table 1. Principles and characteristics of three types of super-resolution microscopy
Principle xy resolution/nm z resolution/nm Fluorescent
moleculesLive cell imaging Image reconstruction STED Stimulated emission depletion based on PSF ~40 ~70 Free Yes Not required SIM Structured illumination ~50 ~250 Free Yes Required PALM Photo activated localization based on
photoswitchable fluorescent molecules~20 ~50 Photoswitchable Yes Required STORM Stochastic optical reconstruction based on
photoswitchable fluorescent molecules~20 ~50 Photoswitchable Yes Required -
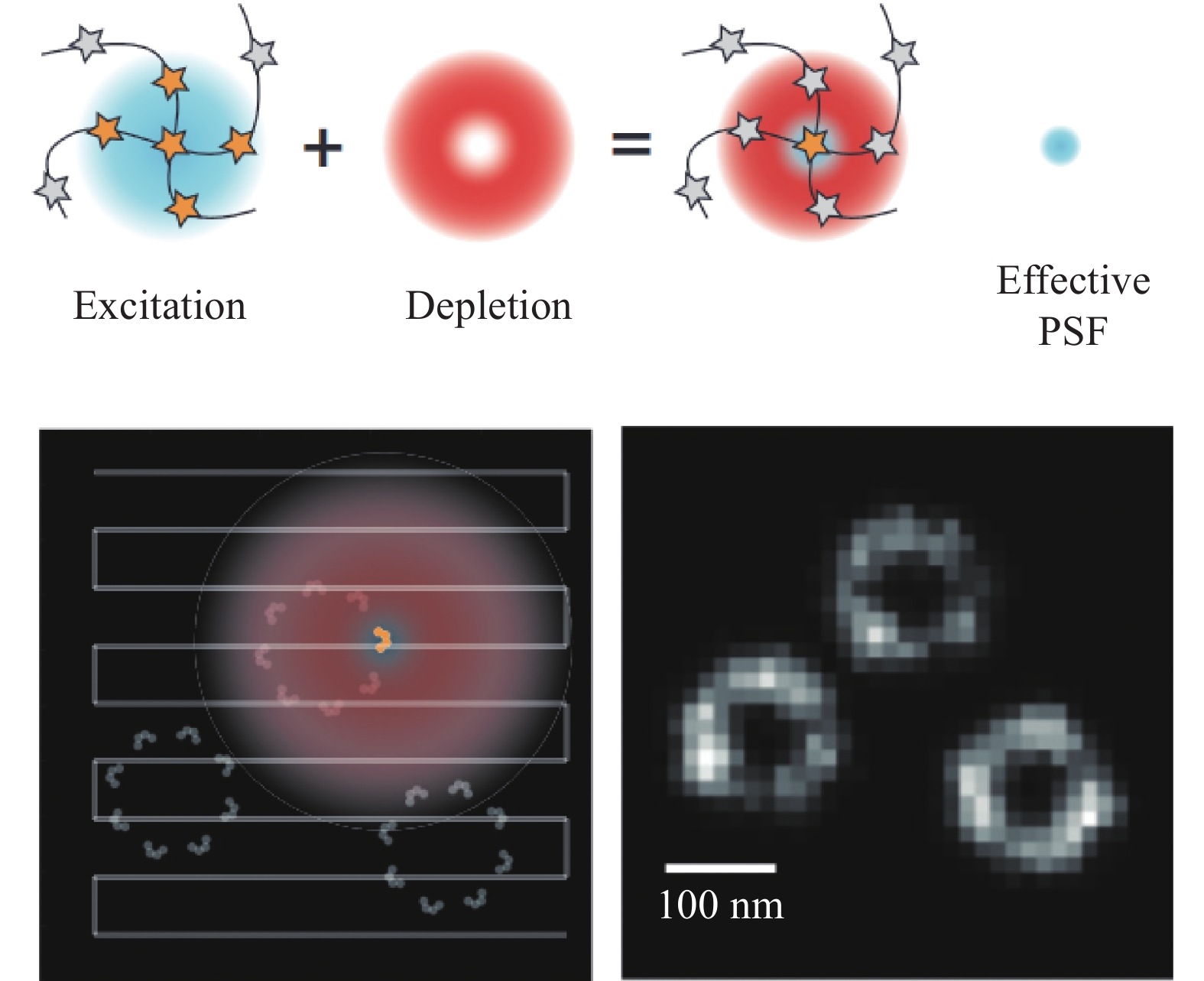
STED显微成像技术以激光共聚焦技术为基础,它通过激发态荧光团的损耗来提高成像分辨率。分辨率取决于激发点的大小,即点扩散函数(Point spread function, PSF),由于PSF内的所有荧光团都会对测量信号产生影响,STED通过引入另一束环形损耗光包围激发光,抑制PSF外围荧光团的信号发射,有效减小激发光斑尺寸,从而提高成像分辨率[16, 20-21](图1)。STED利用纯光学的方法实现超分辨成像,因此它不需要对图像进行后续处理。同时,其分辨率相对较高,可达到40 nm左右的横向分辨率以及70 nm左右的纵向分辨率,而且它的成像速度相对较快,对荧光染料的特异性要求不高[22]。基于以上特点,STED已成为细胞及细胞器精细结构研究中的理想选择之一。但是,STED采用逐点扫描的采集方式导致其时间分辨率仍不能满足活跃的细胞活动,如高度动态的细胞器相互作用过程的成像,这也就限制了其在细胞器相互作用研究中的应用。
-
SIM采用结构化的条纹图案进行照明,通过莫尔效应使传统荧光显微镜无法探测到的样品高频信息进入成像系统的频率探测范围内,实现超分辨成像[23-26](图2)。最初,SIM基于宽场结构光照明,受线性荧光激发特性的影响,其分辨率较宽场成像只能提高两倍,并不能无限制提高。随着非线性荧光激发技术的发展,SIM的横向分辨率获得显著提升,可达到50 nm。虽然宽场结构光照明成像速度快,视场大,但其在厚组织样本中应用受限。近来的研究中,点扫描结构光照明技术使SIM的成像深度有所提升,同时得益于硬件设备的升级和去卷积算法的优化,SIM在保持分辨率的前提下有效提升了成像速度。点扫描SIM还可以与双光子激发相结合,实现厚组织的高分辨率观察[27]。由于SIM具有良好的荧光探针适用性、较高的成像分辨率以及较快的成像速度,已成为活细胞观察中使用最广泛的超分辨成像方法[28-29]。
-
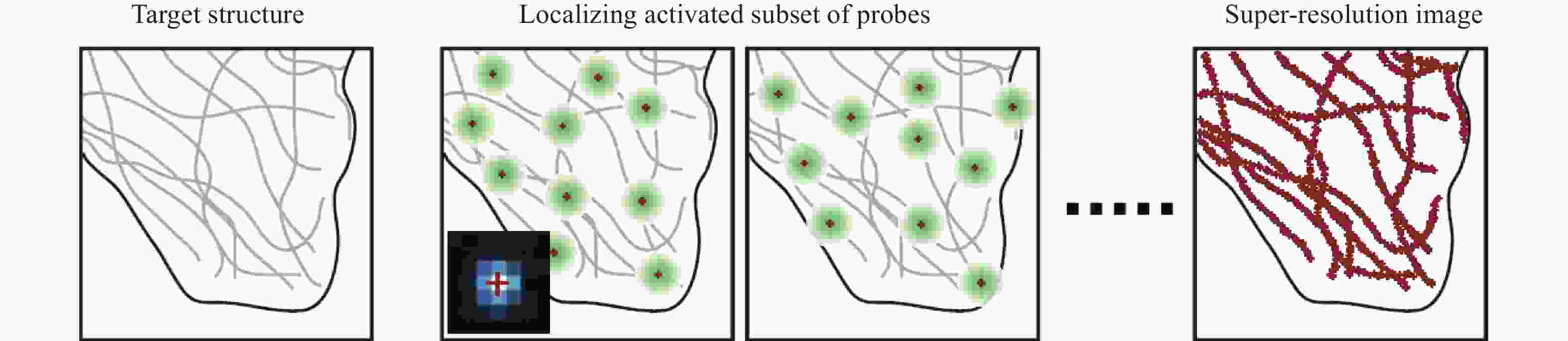
SMLM使用特定的荧光探针,特定波长的激光来激活荧光分子,然后用另一波长激发荧光成像,通过控制激光强度使少量荧光分子激活,采集大量图像后利用单分子定位算法定位单分子的中心位置,并叠加重建得到超分辨图像[18, 30-31](图3),其典型的代表是PALM,可达到10~80 nm的横向分辨率。此外,基于单分子定位技术,庄小威等[32]提出了STORM,通过荧光分子的on-off开关状态采集信号并获得超分辨图像,可实现20~50 nm的横向分辨率。单分子定位显微成像技术已广泛用于组织和细胞成像研究,在细胞器精细结构、蛋白分子的运动等观察中优势明显,它可以呈现微管、线粒体、脂滴甚至核孔复合体等的超微结构,具有观察装置简单、操作方便、分辨率高等优势,但其成像速度低,在活细胞成像中不具优势,且对荧光分子特异性要求较高[33]。
-
超分辨显微成像可以通过多种方式为生物样品提供具有高分辨率结构信息的图像。由于细胞器的结构、形态特征以及细胞器间相互作用中接触位点的形态变化往往小于衍射极限,因此,超分辨显微成像成为研究细胞器的关键技术,在细胞器相互作用研究中具有重要作用。目前,在生物医学领域,超分辨显微成像技术已应用于细胞器精细结构及细胞器间相互作用规律的研究,助力学者从细胞器层面阐明生理病理现象背后的作用机制。
-
线粒体是细胞内主要进行能量供应的细胞器,光学显微镜下呈现棒状、椭圆形或球形等,透射电镜下呈现典型的双层膜结构,嵴清晰可见。直径和长度分别在0.5~1.0 μm和0.5~10 μm的范围内变化[34]。根据细胞功能及能量需求的不同,它的数量也呈动态变化,通常功能旺盛区分布较多,对细胞功能发挥具有重要的作用;溶酶体是细胞内主要进行消化的场所,富含多种水解酶类,常与细胞凋亡、死亡及自噬功能密切相关。光学显微镜及透射电镜下,溶酶体多呈球形,直径约为0.2~0.8 µm。线粒体与溶酶体不仅具有独立的功能,而且两者间存在相互作用来调控细胞功能。以往的研究中借助分子生物学方法可从细胞水平研究线粒体与溶酶体相互作用(主要为线粒体自噬),借助激光共聚焦显微镜观察到线粒体与溶酶体的相互作用主要是受损线粒体通过与溶酶体的融合[35],或受损线粒体产生的囊泡与溶酶体融合来实现线粒体自噬的发生。然而,传统方法并不能观测到更加精细的结构,无法为线粒体与溶酶体的相互作用观察提供足够的分辨率。超分辨显微成像技术的出现,为研究线粒体与溶酶体相互作用规律提供了新手段。
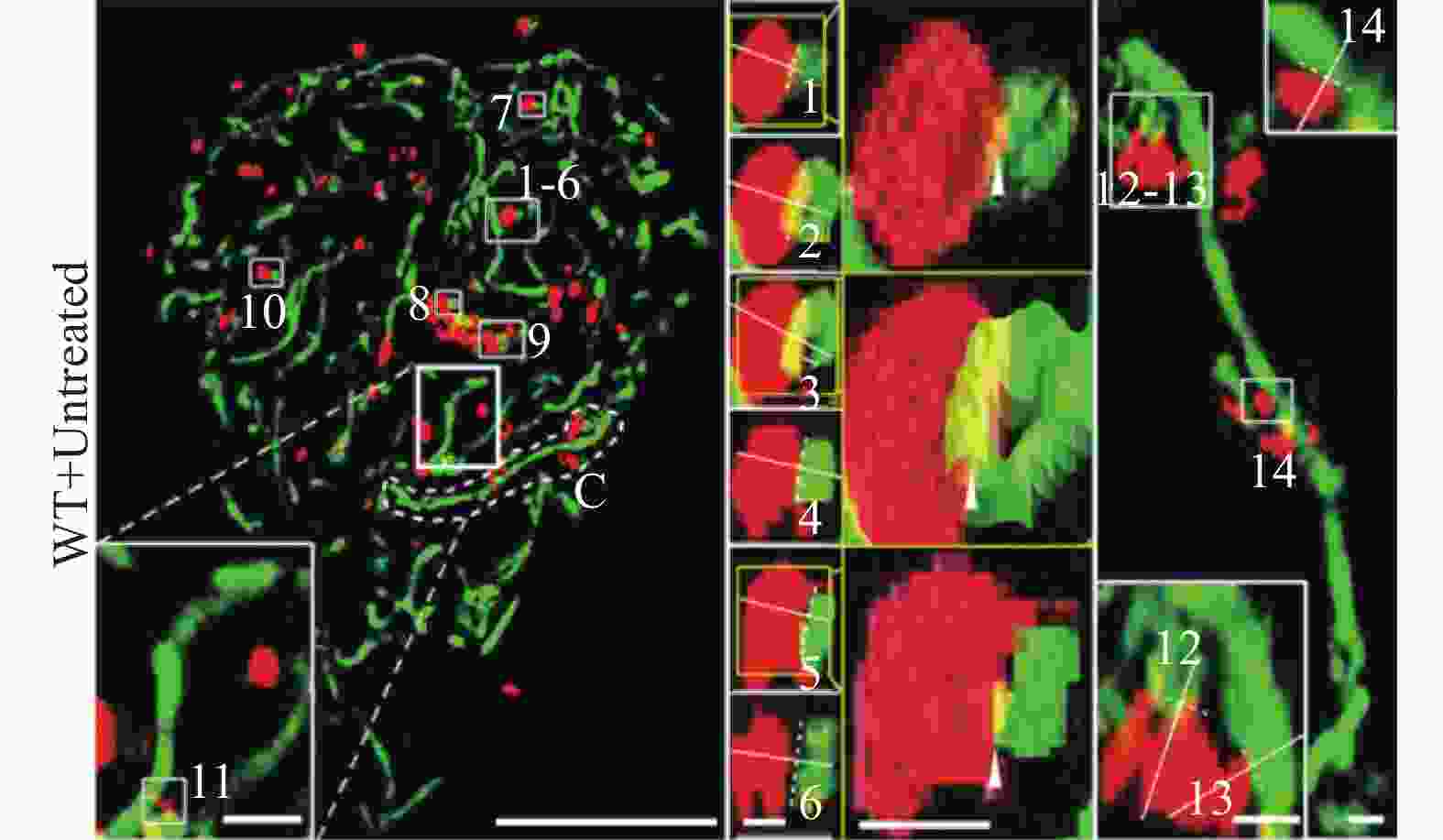
目前研究发现的线粒体与溶酶体相互作用主要包括:正常或病理条件下线粒体-溶酶体接触(Mitochondria and lysosome contact, MLC)和线粒体自噬。Wong等通过N-SIM (尼康,100倍油镜,NA1.49)观察发现,MLC可标记线粒体的分裂位置,并且MLC的连接与连接的解除受到线粒体与溶酶体的双向调控,有可能为揭示人类疾病中这两个细胞器的共同功能障碍提供一定的理论支持[36]。Kim P K等通过SIM观察(蔡司Elyra PS.1,63倍油镜,NA1.4,曝光时间为75 ms,激光功率在3%~10%之间变化,增益水平在60~80之间),发现内质网可招募溶酶体到达线粒体分裂部位,进而协同调节线粒体的分裂[37]。同时,有研究报道线粒体与溶酶体相互作用的分析方法,通过SIM(OMX 3D-SIM,奥林巴斯100倍油镜,NA1.49,最大分辨率横向120 nm,纵向340 nm)可清晰地捕获各种线粒体与溶酶体的相互作用形式,分析得到M值(公式计算所得指标),MLC中M值小于0.4,线粒体自噬中的M值在0.5~1.0之间,该方法为定量分析高分辨率成像下的线粒体-溶酶体相互作用提供了更加简便的判断依据(图4)[38]。此外,研究人员结合SIM (OMX 3D-SIM,60倍油镜,NA1.42)开发了一种溶酶体靶向生物传感器,通过观察溶酶体形态和荧光信号变化来检测线粒体的形态变化与病理条件下的MLC现象。研究发现:溶酶体应激条件下,线粒体的形态从长条状变为圆球形,这为跟踪生理和病理下线粒体-溶酶体相互作用提供了新手段[39]。
-
细胞核是由染色质、RNA、蛋白质组成的动态网状结构,对基因组的折叠和调控至关重要,是真核生物细胞遗传与代谢的重要调控中心[40]。高等动物中细胞核的直径约为5~10 μm,二维观察呈圆形或卵圆形,三维重建后呈不规则球形。线粒体是细胞的能量代谢中心,它的功能受细胞核活动的严格调控,两者之间的相互作用研究对理解它们在生理病理过程中的功能具有重要意义。
近年来,有学者利用透射电镜以及高分辨率激光共聚焦显微镜发现了线粒体与细胞核的特定接触区域,结合分子生物学手段发现核膜上的Cnm1蛋白与线粒体外膜上的Tom70相互作用来介导两者的接触[41]。此外,还有学者发现线粒体与核膜的接触促进线粒体与细胞核的逆行通讯,来驱动细胞对压力的适应[42]。目前,新出现的自我补体分裂荧光蛋白(FP)以及开发的三方探针mito-STAR技术能够观察到细胞核与线粒体之间的物质交换。利用SIM观察(尼康100倍油镜,NA1.49,Andor iXon3 EMCCD),该技术能够用于检测可视化Sirt4蛋白在单细胞中线粒体和细胞核之间的分布,证明Sirt4参与线粒体到细胞核的逆行信号传递[43-44]。以SIM为代表的超分辨显微成像技术呈现了线粒体与细胞核的相互接触作用中更加清晰的细节,但是现有研究中,线粒体与细胞核相互作用的观察手段仍主要集中于透射电镜和共聚焦显微镜。相信随着超分辨显微成像技术的普及,将会揭示更多生理病理过程中两者之间的相互作用规律。
-
细胞骨架是真核细胞中的蛋白纤维网络结构,主要由微丝、微管和中间丝组成,为细胞及细胞器提供结构支持。它具有复杂的功能,包括调节细胞形状和大小、参与细胞分裂、迁移、细胞-细胞相互作用、蛋白质摄取和分泌、受体信号传导等生理活动[45]。构成细胞骨架的微丝由肌动蛋白和肌球蛋白组成,直径约为4~7 nm,主要位于细胞表面,决定细胞的形状并负责细胞的运动;微管由α、β及γ微管蛋白构成,直径约为22~25 nm,它主要决定膜性细胞器的位置,并作为膜泡运输的轨道;中间丝,直径介于微管和微丝之间,约为8~10 nm,它使细胞具有张力和抗剪切力[46]。
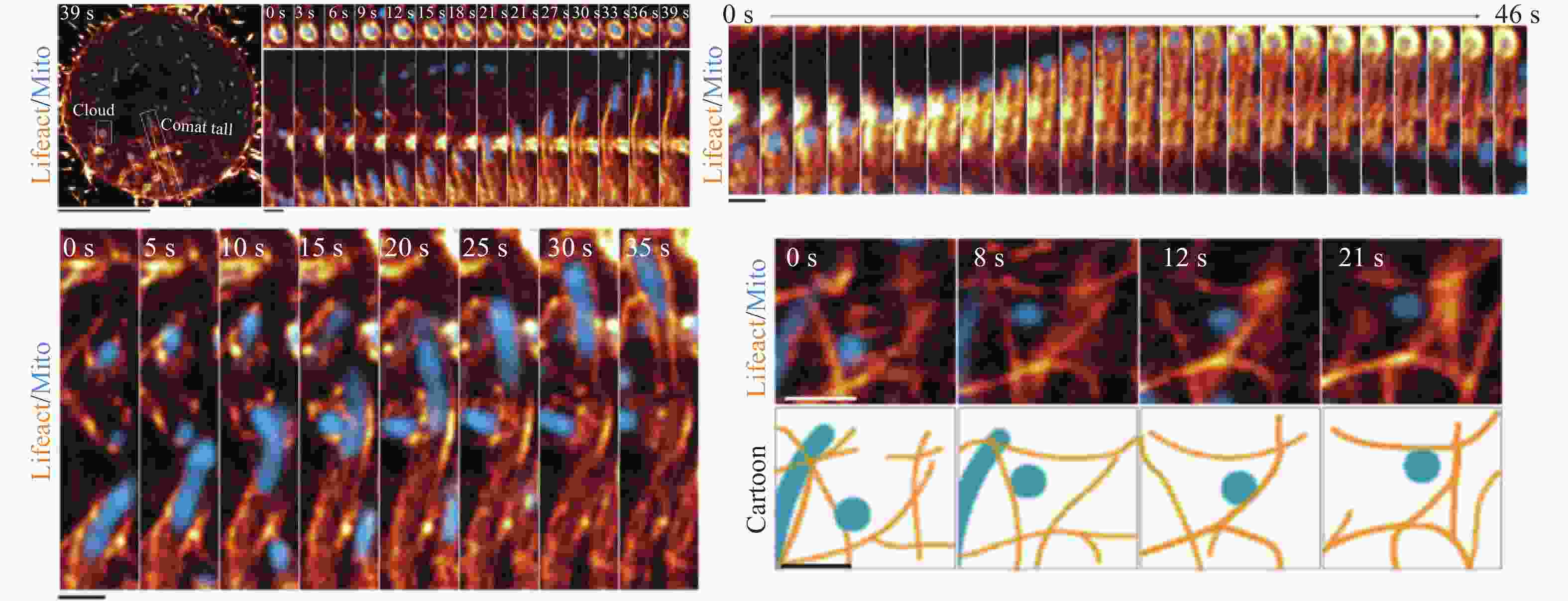
细胞骨架(微管、中间丝、微丝)在维持线粒体的形状、位置和运动方面起着核心作用[47]。线粒体可以沿着微管进行两个方向的移动:顺行(沿微管朝向细胞膜方向移动)和逆行(沿微管朝向细胞核方向移动)。近来的研究中,利用GI-SIM (奥林巴斯100倍物镜,NA1.49,滨松ORCA-Flash 4.0 sCMOS相机)观察发现,在有丝分裂过程中,由肌动蛋白构成的网络支架结构可以对线粒体进行三维定位,肌动蛋白在线粒体表面进行可逆聚集,确保线粒体在对称分裂中的平等和随机遗传[48](图5)。此外,Congying Wu等通过SIM(尼康100倍油镜,NA1.49)观察发现,分支肌动蛋白可以影响微管的乙酰化水平,进而控制细胞内线粒体的分布[49]。细胞骨架的形态同样可以影响线粒体的分裂融合,长微管的突变细胞内线粒体较少,而短微管的细胞内线粒体倾向于分裂,数目增多[50]。线粒体是信号通路的枢纽之一,它的功能改变也可以影响细胞骨架中肌动蛋白调节蛋白的活性,从而调控包括细胞迁移等在内的细胞行为。最近的研究中,冷冻电镜揭示了线粒体与细胞骨架锚定部位的超微结构,超分辨显微镜则将在两者动态互作研究中发挥更重要的作用。
-
脂滴(Lipid droplets, LDs)在部分原核生物以及所有真核生物中均存在,它是细胞内中性脂的主要贮存场所,可以通过膜表面的接触位点与其他细胞器相互作用,参与脂质的储存和分解。同时,它还参与脂肪酸转运、蛋白储存及降解、炎症反应、病毒复制等细胞活动,协同调控细胞内一系列生理生化过程[51-52]。
在不同的细胞内,脂滴的大小差异较大,直径范围约为0.05~200.00 μm[53]。利用透射电镜观察到脂滴的结构基本相同,是一种呈球状的细胞器,均以中性脂为内核,表面覆盖磷脂单分子层及功能蛋白[54]。 激光扫描共聚焦显微镜是目前脂滴荧光成像使用最广泛的显微镜,有多种荧光探针可以标记脂滴。而随着超分辨显微成像技术的发展,对脂滴的动态过程及其与细胞器的相互作用研究变得更加深入。研究者利用超分辨显微成像观察到,与细胞脂质代谢相关的围脂滴蛋白5(Plin5)在线粒体和脂滴相互作用中扮演着重要角色,即过表达 Plin5后脂滴周围的线粒体数目明显增多。利用STED (徕卡,型号TCS SP8,100倍物镜,NA1.4)成像,研究者发现,在脂肪分解过程中,Plin5除了定位在细胞质中,还存在于线粒体附近以及线粒体和脂滴的相互作用位点(图6),并且Plin5的过表达不仅会将线粒体招募至脂滴附近,还会导致出现更小的线粒体结构,且它们会部分或完全包围单个或整个脂滴集群[55-56]。由此,研究者们利用STED发现Plin5在脂滴和线粒体接触部位表达丰富,并在能量需求增加时促进脂滴转运脂肪酸至线粒体,以促进线粒体脂肪氧化。
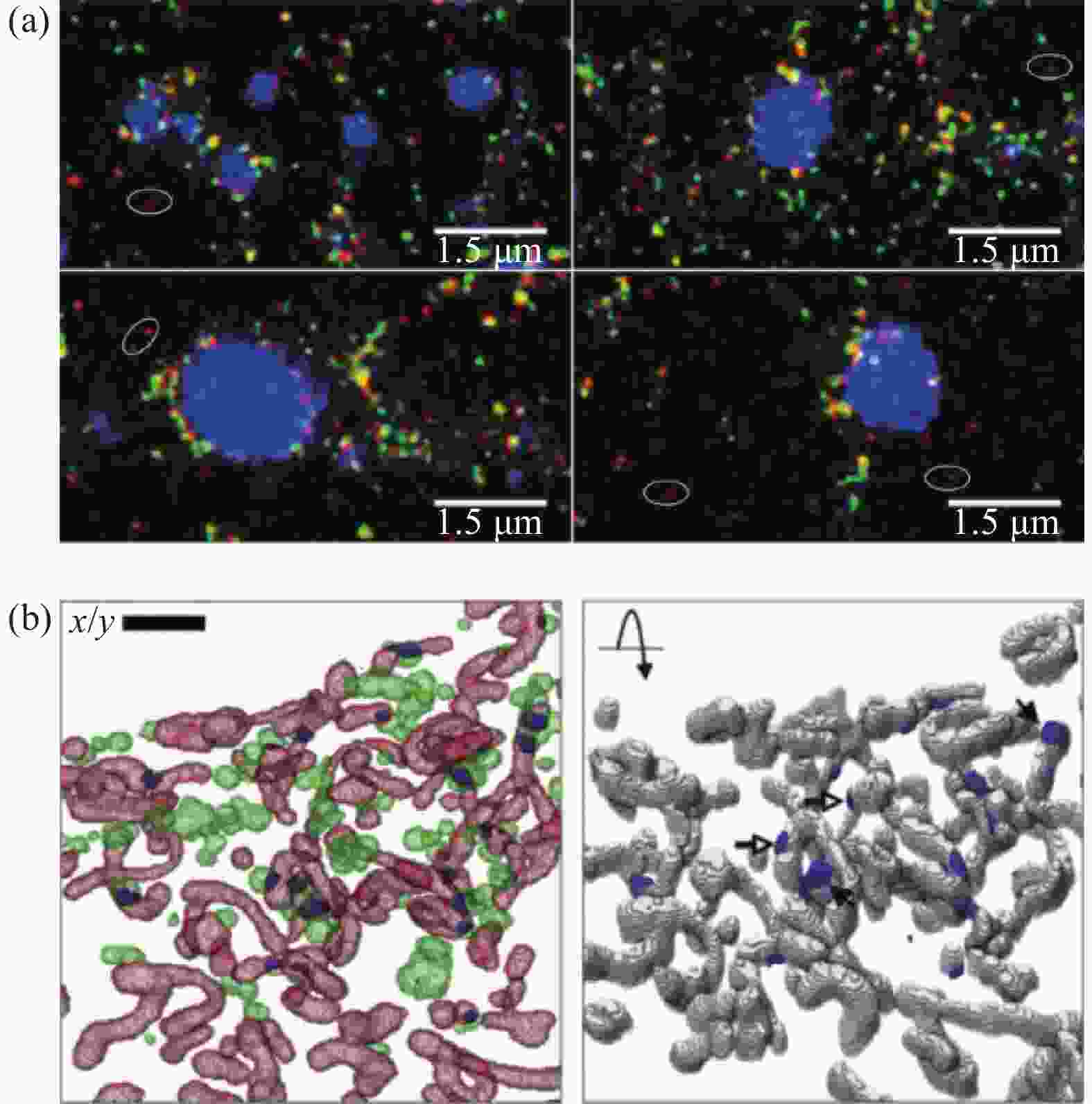
图 6 STED观察线粒体-脂滴接触。(a) 接触面PLIN5的表达;(b) 线粒体-脂滴接触区渲染图(绿色:脂滴;红色:线粒体;蓝色:接触区)[55-56]
Figure 6. STED showed the mitochondrion-lipid droplet interface. (a) Expression of PLIN5 (red) at interface of mitochondria and lipid droplet; (b) Voxelized rendering of extracted LD, mitochondria and overlapping zones between them (LD (green), mitochondria (red) and computed contact zones (blue))[55-56]
-
内质网(ER)是细胞内最大的亚细胞器,约占细胞质体积的35%,呈网络状结构,在脂质和蛋白质生物合成中具有重要的作用[57-58]。内质网在光镜下首次被观察到时,被称为“基底丝”[59]。随着电镜技术的发展,内质网的细微结构被渐渐发现。电镜下内质网管腔有脂质双层结构,呈片状和管状,小管直径约80~100 nm[60-62],而片状呈双层扁平结构,双层间距约为30~50 nm[63]。近来高分辨率观察显示,一些最初被认为是片状结构实际上由致密的基质小管组成[62-64]。
内质网与线粒体(ER-Mito)具有广泛的相互接触,这对包括线粒体分裂、钙信号传导、脂质转运、和线粒体DNA合成在内的多种功能的实现具有重要的作用[10, 65]。因此,观察并揭示ER-Mito相互作用机制具有重要意义,而超分辨显微成像技术的应用则为ER-Mito相互接触和作用的研究提供了直接证据。Dong Li等搭建了掠入射结构光照明显微镜(GI-SIM),利用NA1.7和1.49的100倍油镜以最高97 nm横向分辨率和266 frame/s时间分辨率,发现ER-Mito接触点标记了线粒体分裂的位置,并且ER-Mito接触点参与线粒体分裂的全程:接触点处的线粒体膜拉长成管状,直到分裂为两个独立的线粒体;同时在线粒体融合过程中,内质网小管的参与可加速线粒体的融合[64, 66](图7)。由此可见,ER-Mito接触可介导线粒体的分裂融合过程。
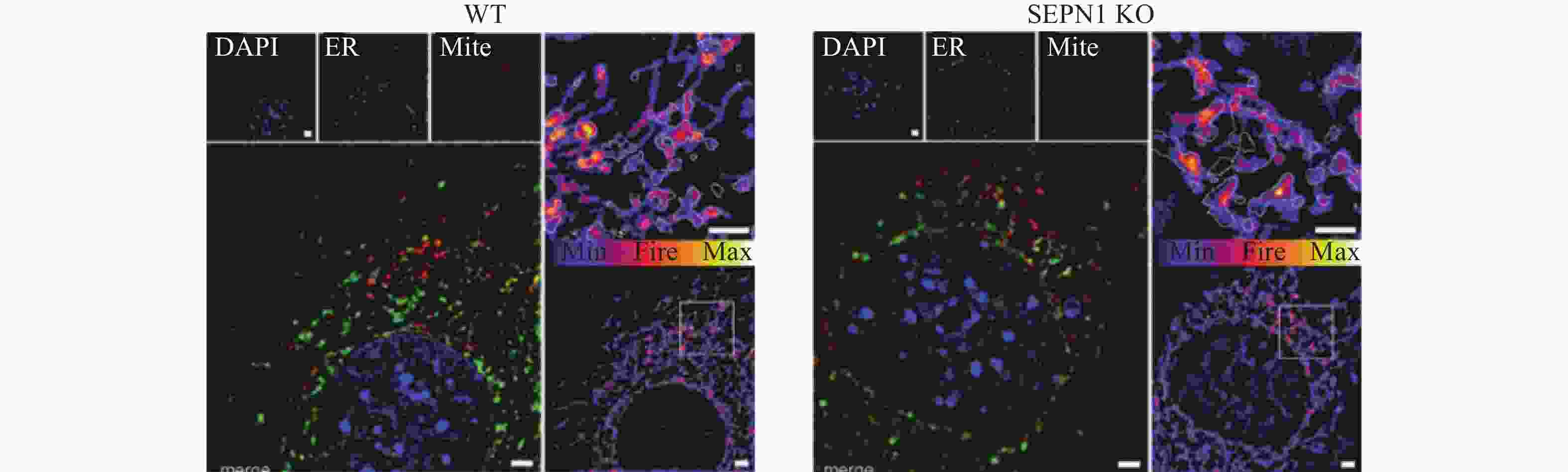
借助SIM (尼康100倍油镜,NA1.49,安道尔iXon3® EMCCD相机,可实现最大横向分辨率120~130 nm)观察,学者们发现内质网与线粒体相关膜(Mitochondria-associated membranes, MAMs)的接近程度与线粒体动力学密切相关,局部Ca2+信号调控线粒体嵴膜(Mitochondrial cristae membranes, CM)动力学和结构,以促进线粒体和内质网之间Ca2+平衡[67]。SIM (尼康100倍油镜,NA1.49)观察发现,内质网和线粒体间会通过MAMs接触并产生钙瞬变。通过特异性追踪线粒体与内质网高分辨率成像,研究SEPN1相关肌肉疾病细胞中的内质网、线粒体和MAMs,发现SEPN1在MAMs中富集,其缺失导致内质网-线粒体接触减少,细胞器Ca2+含量降低,氧化磷酸化过程受损(图8)[68]。
-
内质网负责绝大多数蛋白的合成和修饰,它由片状和管状结构共同形成处于高度动态的膜系统。膜系统的变化主要依赖于管状结构的延伸与收缩,并直接决定内质网的分布和形态。随着成像技术的发展,学者们发现溶酶体和内质网存在协同运动。通过GI-SIM成像,Dong Li等发现35%内质网管状延伸次数与溶酶体的协同运动相关[64],随后Clemens等通过高分辨率成像并构建内质网卷积神经网络,发现溶酶体与内质网偶联,它对内质网小管伸长的贡献率约为73%左右,证实了溶酶体对内质网动态结构的实时调控[15]。利用双色溶酶体单粒子追踪同时跟踪溶酶体和内质网,发现几乎所有(98%)溶酶体都与局部内质网同步运动,尤其是在内质网延长的小管尖端附近。因此,溶酶体与内质网协同作用,动态调控内质网的形态和分布,是内质网动态变化的动力基础。
另外,也有研究报道,内质网在运输和控制溶酶体方面发挥着重要作用[69]。通过SIM观察,发现内质网小管可以收缩并紧密围绕溶酶体,当这些被包围的溶酶体接近微管时会倾向于发生运动,且在运动期间保持内质网和移动的溶酶体前缘之间的接触,促使溶酶体的长距离运输[64]。
-
依赖于细胞微管(MT)的内质网(ER)小管运动是ER结构和功能的基础。以往的研究发现,ER与MT通过滑动机制和尖端连接复合物机制可实现ER小管的形成和运动。在真核细胞中,虽已知ER和MT会经历高度动态但有组织的相互作用,协调复杂的细胞功能,但由于之前技术手段的限制,无法进行更精细化的研究。随着超分辨显微成像技术的出现及应用,使得对ER和MT相关作用功能的研究可以更深入展开。
借助GI-SIM (NA1.7和NA1.49的100倍油镜,横向分辨率最高97 nm,时间分辨率266 frame/s)发现,哺乳动物细胞中新的ER小管通过“搭便车”的方式锚定在移动的微管(MT)上,随之被拉长,并与现有ER小管融合形成新的三向连接[64]。此外,有研究通过3D-SIM (DeltaVision,OMX Blaze,100倍物镜,横向分辨率120 nm,纵向分辨率340 nm)观察,发现了基质相互作用分子1 (STIM1)-MTs-ER相互作用模型。在神经元生长锥中钙信号的时空调节对于轴突引导至关重要,借助STIM1和动态末端结合蛋白3可以将ER偶联到MTs上,使得MTs和ER衍生的钙信号相结合,为钙信号在新生神经中引导生长锥的空间定位提供了一个新模型[70]。此外,STED成像结合计算模拟发现,ER小管连接密度的空间不均匀会导致作用于MT的整体收缩力,并影响它们的分布,揭示了内质网动力学在细胞内微管分布中起着重要作用[15]。
-
超分辨显微成像技术突破了光学衍射极限,使活细胞及其细胞器成像分辨率有了显著的提升,为从细胞器层面研究细胞功能提供了新手段。通过总结现有的部分研究发现(表2),结构光照明显微成像(SIM)在细胞器相互作用研究中更受青睐,其次是受激发射损耗显微成像(STED),而单分子定位显微成像(SMLM)在活细胞中细胞器相互作用研究中应用少见,可能与其时间分辨率不足有关。
表 2 超分辨显微成像技术在细胞器相互作用研究中的应用
Table 2. Application of super-resolution microscopy in the study of organelle interactions
Organelle interactions Research contents Super-resolution microscopy Cells Mitochondria-lysosome Mitochondria and lysosome contact ( MLC) marks sites of mitochondrial fission and the formation and release of MLC is bidirectionally regulated by mitochondrial and lysosomal dynamic [36]; Endoplasmic reticulum (ER) recruits lysosomes to act in concert at the fission site for the efficient division of mitochondria [37]; The analysis methods of mitochondria-lysosome interactions based on structured illumination microscopy
(SIM) [38]; Lysosome-targeted biosensor for the super-resolution imaging of lysosome-mitochondrion interaction based on SIM[39].SIM HT22 cells; SHSY-5Y cells; HeLa cells; HSF cells Mitochondria- nucleus Observation of mitochondria-nucleus contact and the change of expression sites of Sirt4 from mitochondria to nucleus under mitochondrial stress conditions [43-44]. SIM HeLa cells; Pancreatic β cells Mitochondria-cytoskeleton Mitochondrial morphology and distribution are regulated by cytoskeleton. During mitosis, dense meshwork of subcortical actin cables organizes three-dimensional mitochondrial positioning to ensure both equal and random inheritance of mitochondria in symmetrically dividing cells [48]; Actin maintains microtubule organization, dynamics and stability by affecting tubulin acetylation levels and further regulate mitochondrial distribution [49]; Cytoskeleton regulates fission and fusion of mitochondria [50]. SIM; STED Hela cells; COS-7 cells; HEK293 cells; U2 OS cells Mitochondria-lipid droplet Observation of mitochondria-lipid droplets contacts; Overexpression of perilipin5 leads to increased number of mitochondria surrounding lipid droplets [55]. STED COS-7 cells Mitochondria-Endoplasmic reticulum Tubular endoplasmic reticulum regulates mitochondrial fission and fusion [64]; Calcium transients on the sites of mitochondria-endoplasmic reticulum contacts[67]; In SEPN1-related myopathy, SEPN1 deficiency results in less mitochondria-endoplasmic reticulum contacts, calcium contents and damaged oxidative phosphorylation process [68]. SIM COS-7 cells; U2 OS cells; HeLa cells Endoplasmic reticulum-lysosome Lysosomes moved synchronously with local endoplasmic reticulum. The anchorage of lysosomes to endoplasmic reticulum growth tips is critical for endoplasmic reticulum tubule elongation and connection[15]; Endoplasmic reticulum contacts with the edge of lysosome, which promotes the long-distance transportation of lysosome[64]. SIM COS-7 cells; U2 OS cells Endoplasmic reticulum-cytoskeleton Endoplasmic reticulum anchors to microtubules, which guides the formation of new endoplasmic reticulum tubule branches[64]; Endoplasmic reticulum dynamics play important roles in microtubules distribution[15]. SIM; STED COS-7 cells; U2 OS cells 超分辨显微成像技术在细胞器及其相互作用研究中的贡献主要体现在以下三方面:获得细胞器结构的直观精细高分辨率图像;为已发现的细胞器相互作用提供了高分辨率图像直接证据;观察到细胞器间相互作用的新形式。超分辨显微成像技术虽然在细胞器精细结构观察与细胞器相互作用研究中具有其他技术不能替代的作用,但由于成像的分辨率、速度和成像深度,以及荧光探针的光毒性等问题(表3),在实际应用中仍存在一些不足,限制了其在该领域内的应用效果。
表 3 超分辨显微成像技术的应用参数
Table 3. Application parameters of super-resolution microscopy
STED SIM SMLM xy resolution/nm ~40 ~50 ~20 z resolution/nm ~70 ~250 ~50 Temporal resolution 5 ms-2 s 10-500 ms 1 min-1 h Light intensity Medium-high Low-medium Low-medium Live cell dynamic imaging Medium Good Poor (1) STED在对活细胞细胞器相互作用研究中,在横向分辨率上突破了衍射极限,但由于该技术采用的是点扫描技术,在对大视野区域进行成像时,图像采集比普通荧光显微镜耗时长,限制了面积较大的活细胞等生物样本内生物分子的动态过程或相互作用的实时成像;同时由于使用高功率的损耗光,产生的光毒性会对生物活性样本产生一定的光损伤,限制了细胞器及相互作用成像的准确性,因此可从提高成像速度和降低损耗光功率两方面改进,进一步提高STED在活细胞中细胞器及相互作用研究中的适用性。
(2) SIM在对活细胞细胞器相互作用研究中,在横向与纵向分辨率上均有所提高,可实现活细胞中细胞器的3D高分辨率成像,同时具有高速扫描、多色成像以及低光漂白和低光毒性等优势,非常适合研究细胞器动力学。与TIRF或TIRF-SIM相比,GI-SIM荧光强度更大;与SDCM相比,它提供了更好的空间分辨率和更快的成像速度;与其他超分辨显微成像技术相比,它使时空分辨率与光漂白和光毒性之间更好地平衡。然而由于只能将分辨率提高到宽场成像两倍,因此提高SIM的成像分辨率可提高其在细胞器相互作用中的应用。
(3)目前单分子定位显微成像SMLM,主要包括光激活定位显微成像(PALM)与随机光学重构显微成像(STORM)。该技术可将分辨率提高到20 nm以下,PALM技术通过细胞自身表达荧光蛋白进行成像,更适合用于活细胞内蛋白的超分辨成像。SMLM在荧光团本身稀疏标记的样本中,可用于活样本中进行超分辨率跟踪,还可实现活体样本的3D成像。但是由于分子定位具有顺序性,需要反复激活-淬灭荧光分子获得许多幅原始图像重构获得超分辨图像,因此其成像的时间分辨率相对较低,限制了其在较为活跃的细胞器相互作用成像中的应用。
超分辨显微成像技术的应用为活细胞中细胞器相互作用研究提供了新手段,然而现有研究中仍存在一些不足,如多细胞相互作用成像受到空间分辨率的限制,动态变化的成像受到时间分辨率的限制,高功率激发光产生的光毒性无法避免对活细胞产生影响等。因此,未来超分辨显微成像技术在活细胞的细胞器相互作用研究应用中,将在提高空间和时间分辨率,降低激发光功率,开发低光毒性荧光染料,降低光毒性对活细胞的影响等方面进一步提升。此外,人工智能算法在超分辨显微成像技术中的应用愈加广泛,如何利人工智能算法来提升成像的效果以及分析细胞器相互作用规律也将是重点研究的方向之一。总之,超分辨显微成像技术的出现为揭示细胞器相互作用规律提供了有力的技术手段,对于探索细胞器互作异常引发疾病背后的机制具有非常重要的意义。
Application of super-resolution microscopy in the study of organelle interactions (invited)
-
摘要: 观察细胞器间动态相互作用,深入分析作用规律,对于揭示生理病理过程现象背后的机制具有十分重要的意义。传统光学显微镜受到由光波波长和孔径造成的衍射极限的限制,无法观测细胞器纳米级精细结构及细胞器间相互作用的动态变化规律。超分辨显微成像技术的出现为细胞器相互作用研究提供了重要手段,在深入揭示细胞器相互作用规律,阐明生理病理现象深层的机制研究中发挥了重要的作用。文中介绍了受激发射损耗(Stimulated emission depletion, STED)显微成像、结构光照明显微成像(Structured illumination microscopy, SIM)、单分子定位显微成像(Single molecule localization microscopy, SMLM)技术,并总结了这三类超分辨显微成像技术在细胞器相互作用中的应用与现状,为超分辨显微成像技术在细胞器相互作用研究中的应用提供思路拓展。最后,对超分辨显微成像技术在细胞器相互作用研究中的优势与不足进行分析总结,展望了超分辨显微成像技术在活细胞内细胞器相互作用成像中的需求发展趋势,为光学与医学及生物学的交叉融合发展提供一定的参考。
-
关键词:
- 超分辨显微成像 /
- 细胞器间相互作用 /
- 受激发射损耗显微成像 /
- 结构光照明显微成像 /
- 单分子定位显微成像
Abstract: Observing dynamic interaction between organelles and analyzing the law of action is of great significance for revealing the mechanism behind the phenomenon of physiological and pathological processes. Due to the limitation of the optical diffraction determined by wavelength and aperture, traditional optical microscopes cannot observe the nanoscale fine structure of organelles and the dynamic changes of interactions among them. The emergence of super-resolution microscopy imaging technology provides an important mean for the study of organelle interaction. This paper introduces the fluorescence microscopy (STED), structured illumination imaging (SIM), and single-molecule localization imaging (SMLM). The application of these super-resolution microscopy in the study of dynamic interaction between organelles provides the expansion of application ideas for super-resolution microscopy. Finally, the advantages and disadvantages of super-resolution microscopy in the study of organelle interactions are analyzed. In conclusion, the demand and development trend of super-resolution microscopy technology in the imaging of intracellular organelle interaction in living cells is prospected, which provides a certain reference for the cross-integration development of optics, medicine and biology. -
图 6 STED观察线粒体-脂滴接触。(a) 接触面PLIN5的表达;(b) 线粒体-脂滴接触区渲染图(绿色:脂滴;红色:线粒体;蓝色:接触区)[55-56]
Figure 6. STED showed the mitochondrion-lipid droplet interface. (a) Expression of PLIN5 (red) at interface of mitochondria and lipid droplet; (b) Voxelized rendering of extracted LD, mitochondria and overlapping zones between them (LD (green), mitochondria (red) and computed contact zones (blue))[55-56]
表 1 三类超分辨显微成像技术的原理及特点
Table 1. Principles and characteristics of three types of super-resolution microscopy
Principle xy resolution/nm z resolution/nm Fluorescent
moleculesLive cell imaging Image reconstruction STED Stimulated emission depletion based on PSF ~40 ~70 Free Yes Not required SIM Structured illumination ~50 ~250 Free Yes Required PALM Photo activated localization based on
photoswitchable fluorescent molecules~20 ~50 Photoswitchable Yes Required STORM Stochastic optical reconstruction based on
photoswitchable fluorescent molecules~20 ~50 Photoswitchable Yes Required 表 2 超分辨显微成像技术在细胞器相互作用研究中的应用
Table 2. Application of super-resolution microscopy in the study of organelle interactions
Organelle interactions Research contents Super-resolution microscopy Cells Mitochondria-lysosome Mitochondria and lysosome contact ( MLC) marks sites of mitochondrial fission and the formation and release of MLC is bidirectionally regulated by mitochondrial and lysosomal dynamic [36]; Endoplasmic reticulum (ER) recruits lysosomes to act in concert at the fission site for the efficient division of mitochondria [37]; The analysis methods of mitochondria-lysosome interactions based on structured illumination microscopy
(SIM) [38]; Lysosome-targeted biosensor for the super-resolution imaging of lysosome-mitochondrion interaction based on SIM[39].SIM HT22 cells; SHSY-5Y cells; HeLa cells; HSF cells Mitochondria- nucleus Observation of mitochondria-nucleus contact and the change of expression sites of Sirt4 from mitochondria to nucleus under mitochondrial stress conditions [43-44]. SIM HeLa cells; Pancreatic β cells Mitochondria-cytoskeleton Mitochondrial morphology and distribution are regulated by cytoskeleton. During mitosis, dense meshwork of subcortical actin cables organizes three-dimensional mitochondrial positioning to ensure both equal and random inheritance of mitochondria in symmetrically dividing cells [48]; Actin maintains microtubule organization, dynamics and stability by affecting tubulin acetylation levels and further regulate mitochondrial distribution [49]; Cytoskeleton regulates fission and fusion of mitochondria [50]. SIM; STED Hela cells; COS-7 cells; HEK293 cells; U2 OS cells Mitochondria-lipid droplet Observation of mitochondria-lipid droplets contacts; Overexpression of perilipin5 leads to increased number of mitochondria surrounding lipid droplets [55]. STED COS-7 cells Mitochondria-Endoplasmic reticulum Tubular endoplasmic reticulum regulates mitochondrial fission and fusion [64]; Calcium transients on the sites of mitochondria-endoplasmic reticulum contacts[67]; In SEPN1-related myopathy, SEPN1 deficiency results in less mitochondria-endoplasmic reticulum contacts, calcium contents and damaged oxidative phosphorylation process [68]. SIM COS-7 cells; U2 OS cells; HeLa cells Endoplasmic reticulum-lysosome Lysosomes moved synchronously with local endoplasmic reticulum. The anchorage of lysosomes to endoplasmic reticulum growth tips is critical for endoplasmic reticulum tubule elongation and connection[15]; Endoplasmic reticulum contacts with the edge of lysosome, which promotes the long-distance transportation of lysosome[64]. SIM COS-7 cells; U2 OS cells Endoplasmic reticulum-cytoskeleton Endoplasmic reticulum anchors to microtubules, which guides the formation of new endoplasmic reticulum tubule branches[64]; Endoplasmic reticulum dynamics play important roles in microtubules distribution[15]. SIM; STED COS-7 cells; U2 OS cells 表 3 超分辨显微成像技术的应用参数
Table 3. Application parameters of super-resolution microscopy
STED SIM SMLM xy resolution/nm ~40 ~50 ~20 z resolution/nm ~70 ~250 ~50 Temporal resolution 5 ms-2 s 10-500 ms 1 min-1 h Light intensity Medium-high Low-medium Low-medium Live cell dynamic imaging Medium Good Poor -
[1] Yang Z, Samanta S, Yan W, et al. Super-resolution microscopy for biological imaging [J]. Adv Exp Med Biol, 2021, 3233: 23-43. [2] Heald R, Cohen-Fix O. Morphology and function of membrane-bound organelles [J]. Curr Opin Cell Biol, 2014, 26: 79-86. doi: 10.1016/j.ceb.2013.10.006 [3] Yang Z, Zhang Z, Zhao Y, et al. Organelle interaction and drug discovery: Towards correlative nanoscopy and molecular dynamics simulation [J]. Front Pharmacol, 2022, 13: 935898. doi: 10.3389/fphar.2022.935898 [4] Lemon W C, Mcdole K. Live-cell imaging in the era of too many microscopes [J]. Curr Opin Cell Biol, 2020, 66: 34-42. doi: 10.1016/j.ceb.2020.04.008 [5] Lboukili I, Stamatas G, Descombes X. Automating reflectance confocal microscopy image analysis for dermatological research: A review [J]. J Biomed Opt, 2022, 27(7): 070902. [6] Bourzac K. Cell imaging: Beyond the limits [J]. Nature, 2015, 526(7574): S50-S54. doi: 10.1038/526S50a [7] Arizono M, Idziak A, Quici F, et al. Getting sharper: The brain under the spotlight of super-resolution microscopy [J]. Trends Cell Biol, 2022, S0962-8924(22): 00150-7. [8] Hu Chunguang, Zha Ridong, Ling Qiuyu, et al. Super-resolution microscopy applications and development in living cell [J]. Infrared and Laser Engineering, 2017, 46(11): 1103002. (in Chinese) doi: 10.3788/IRLA201746.1103002 [9] Lu M, Ward E, van Tartwijk F W, et al. Advances in the study of organelle interactions and their role in neurodegenerative diseases enabled by super-resolution microscopy [J]. Neurobiol Dis, 2021, 159: 105475. doi: 10.1016/j.nbd.2021.105475 [10] Friedman J R, Lackner L L, West M, et al. ER tubules mark sites of mitochondrial division [J]. Science, 2011, 334(6054): 358-362. doi: 10.1126/science.1207385 [11] Rowland A A, Chitwood P J, Phillips M J, et al. ER contact sites define the position and timing of endosome fission [J]. Cell, 2014, 159(5): 1027-1041. doi: 10.1016/j.cell.2014.10.023 [12] Lee J E, Cathey P I, Wu H, et al. Endoplasmic reticulum contact sites regulate the dynamics of membraneless organelles [J]. Science, 2020, 367(6477): eaay7108. doi: 10.1126/science.aay7108 [13] Daniele T, Schiaffino M V. Organelle biogenesis and interorganellar connections: Better in contact than in isolation [J]. Commun Integr Biol, 2014, 7: e29587. doi: 10.4161/cib.29587 [14] Peng W, Wong Y C, Krainc D. Mitochondria-lysosome contacts regulate mitochondrial Ca2+ dynamics via lysosomal TRPML1 [J]. Proc Natl Acad Sci U S A, 2020, 117(32): 19266-19275. doi: 10.1073/pnas.2003236117 [15] Lu M, van Tartwijk F W, Lin J Q, et al. The structure and global distribution of the endoplasmic reticulum network are actively regulated by lysosomes [J]. Sci Adv, 2020, 6(51): eabc7209. doi: 10.1126/sciadv.abc7209 [16] Hell S W, Wichmann J. Breaking the diffraction resolution limit by stimulated emission: Stimulated-emission-depletion fluorescence microscopy [J]. Opt Lett, 1994, 19(11): 780-782. doi: 10.1364/OL.19.000780 [17] Gustafsson M G. Surpassing the lateral resolution limit by a factor of two using structured illumination microscopy[J]. J Microsc, 2000, 198(Pt2): 82-87. [18] Betzig E, Patterson G H, Sougrat R, et al. Imaging intracellular fluorescent proteins at nanometer resolution [J]. Science, 2006, 313(5793): 1642-1645. doi: 10.1126/science.1127344 [19] Sigal Y M, Zhou R, Zhuang X. Visualizing and discovering cellular structures with super-resolution microscopy [J]. Science, 2018, 361(6405): 880-887. [20] Klar T A, Hell S W. Subdiffraction resolution in far-field fluorescence microscopy [J]. Opt Lett, 1999, 24(14): 954-956. doi: 10.1364/OL.24.000954 [21] Liu S, Hoess P, Ries J. Super-resolution microscopy for structural cell biology [J]. Annu Rev Biophys, 2022, 51: 301-326. doi: 10.1146/annurev-biophys-102521-112912 [22] Zhou Hanqiu, Zhu Yinru, Han Hongyi, et al. Research progress of live cell and in vivo super-resolution imaging based on STED [J]. Progress in Biochemistry and Biophysics, 2022, 49: 1-20. (in Chinese) doi: 10.16476/j.pibb.2022.0272 [23] Gustafsson M G. Nonlinear structured-illumination microscopy: wide-field fluorescence imaging with theoretically unlimited resolution [J]. Proc Natl Acad Sci U S A, 2005, 102(37): 13081-13086. doi: 10.1073/pnas.0406877102 [24] Rego E H, Shao L, Macklin J J, et al. Nonlinear structured-illumination microscopy with a photoswitchable protein reveals cellular structures at 50-nm resolution [J]. Proc Natl Acad Sci U S A, 2012, 109(3): E135-E143. [25] Ströhl F, Kaminski C F. Frontiers in structured illumination microscopy [J]. Optica, 2016, 3(6): 667. doi: 10.1364/OPTICA.3.000667 [26] Valli J, Garcia-Burgos A, Rooney L M, et al. Seeing beyond the limit: A guide to choosing the right super-resolution microscopy technique [J]. J Biol Chem, 2021, 297(1): 100791. doi: 10.1016/j.jbc.2021.100791 [27] Chen Ting-ai, Chen Long-chao, et al. Structured illumination super-resolution microscopy technology: review and prospect [J]. Chinese Optics, 2018, 11(3): 307-328. (in Chinese) doi: 10.3788/co.20181103.0307 [28] Castello M, Sheppard C J, Diaspro A, et al. Image scanning microscopy with a quadrant detector [J]. Opt Lett, 2015, 40(22): 5355-5358. doi: 10.1364/OL.40.005355 [29] Sheppard C J, Mehta S B, Heintzmann R. Superresolution by image scanning microscopy using pixel reassignment [J]. Opt Lett, 2013, 38(15): 2889-2892. doi: 10.1364/OL.38.002889 [30] Sauer M, Heilemann M. Single-molecule localization microscopy in eukaryotes [J]. Chem Rev, 2017, 117(11): 7478-7509. [31] Huang B, Bates M, Zhuang X. Super-resolution fluorescence microscopy [J]. Annu Rev Biochem, 2009, 78: 993-1016. doi: 10.1146/annurev.biochem.77.061906.092014 [32] Rust M J, Bates M, Zhuang X. Sub-diffraction-limit imaging by stochastic optical reconstruction microscopy (STORM) [J]. Nat Methods, 2006, 3(10): 793-795. doi: 10.1038/nmeth929 [33] An Sha, Dan Dan, Yu Xiang-hua, et al. Progress and prospect of research on single-molecule localization super-resolution microscopy (Invited Review) [J]. Acta Photonica Sinica, 2020, 49(9): 0918001. (in Chinese) [34] Caicedo A, Aponte P M, Cabrera F, et al. Artificial mitochondria transfer: Current challenges, advances, and future applications [J]. Stem Cells Int, 2017, 2017: 7610414. [35] Wang Y, Li L, Hou C, et al. SNARE-mediated membrane fusion in autophagy [J]. Semin Cell Dev Biol, 2016, 60: 97-104. doi: 10.1016/j.semcdb.2016.07.009 [36] Wong Y C, Ysselstein D, Krainc D. Mitochondria-lysosome contacts regulate mitochondrial fission via RAB7GTP hydrolysis [J]. Nature, 2018, 554(7692): 382-386. [37] Boutry M, Kim P K. ORP1L mediated PI(4)P signaling at ER-lysosome-mitochondrion three-way contact contributes to mitochondrial division [J]. Nat Commun, 2021, 12(1): 5354. doi: 10.1038/s41467-021-25621-4 [38] Chen Q, Shao X, Hao M, et al. Quantitative analysis of interactive behavior of mitochondria and lysosomes using structured illumination microscopy [J]. Biomaterials, 2020, 250: 120059. doi: 10.1016/j.biomaterials.2020.120059 [39] Wang H, Fang G, Chen H, et al. Lysosome-targeted biosensor for the super-resolution imaging of lysosome-mitochondrion interaction [J]. Front Pharmacol, 2022, 13: 865173. doi: 10.3389/fphar.2022.865173 [40] Maruyama D, Ohtsu M, Higashiyama T. Cell fusion and nuclear fusion in plants [J]. Semin Cell Dev Biol, 2016, 60: 127-135. doi: 10.1016/j.semcdb.2016.07.024 [41] Eisenberg-Bord M, Zung N, Collado J, et al. Cnm1 mediates nucleus-mitochondria contact site formation in response to phospholipid levels [J]. J Cell Biol, 2021, 220(11): e202104100. doi: 10.1083/jcb.202104100 [42] Desai R, East D A, Hardy L, et al. Mitochondria form contact sites with the nucleus to couple prosurvival retrograde response [J]. Sci Adv, 2020, 6(51): eabc9955. doi: 10.1126/sciadv.abc9955 [43] Michishita E, Park J Y, Burneskis J M, et al. Evolutionarily conserved and nonconserved cellular localizations and functions of human SIRT proteins [J]. Mol Biol Cell, 2005, 16(10): 4623-4635. doi: 10.1091/mbc.e05-01-0033 [44] Ramadani-Muja J, Gottschalk B, Pfeil K, et al. Visualization of sirtuin 4 distribution between mitochondria and the nucleus, Based on bimolecular fluorescence self-complementation [J]. Cells, 2019, 8(12): 1583. doi: 10.3390/cells8121583 [45] Ivanov A I, Le H T, Naydenov N G, et al. Novel functions of the septin cytoskeleton: Shaping Up tissue inflammation and fibrosis [J]. Am J Pathol, 2021, 191(1): 40-51. doi: 10.1016/j.ajpath.2020.09.007 [46] Weber K, Osborn M. Cytoskeleton: definition, structure and gene regulation [J]. Pathol Res Pract, 1982, 175(2-3): 128-145. doi: 10.1016/S0344-0338(82)80104-0 [47] Kuznetsov A V, Javadov S, Grimm M, et al. Crosstalk between mitochondria and cytoskeleton in cardiac cells [J]. Cells, 2020, 9(1): 222. doi: 10.3390/cells9010222 [48] Moore A S, Coscia S M, Simpson C L, et al. Actin cables and comet tails organize mitochondrial networks in mitosis [J]. Nature, 2021, 591(7851): 659-664. doi: 10.1038/s41586-021-03309-5 [49] Shi P, Wang Y, Huang Y, et al. Arp2/3-branched actin regulates microtubule acetylation levels and affects mitochondrial distribution [J]. J Cell Sci, 2019, 132(6): jcs226506. [50] Mehta K, Chacko L A, Chug M K, et al. Association of mitochondria with microtubules inhibits mitochondrial fission by precluding assembly of the fission protein Dnm1 [J]. J Biol Chem, 2019, 294(10): 3385-3396. doi: 10.1074/jbc.RA118.006799 [51] Rambold A S, Cohen S, Lippincott-Schwartz J. Fatty acid trafficking in starved cells: Regulation by lipid droplet lipolysis, autophagy, and mitochondrial fusion dynamics [J]. Dev Cell, 2015, 32(6): 678-692. doi: 10.1016/j.devcel.2015.01.029 [52] Li Z, Thiel K, Thul P J, et al. Lipid droplets control the maternal histone supply of Drosophila embryos [J]. Curr Biol, 2012, 22(22): 2104-2113. doi: 10.1016/j.cub.2012.09.018 [53] Murphy D J. The biogenesis and functions of lipid bodies in animals, plants and microorganisms [J]. Prog Lipid Res, 2001, 40(5): 325-438. doi: 10.1016/S0163-7827(01)00013-3 [54] Walther T C, Chung J, Farese R J. Lipid droplet biogenesis [J]. Annu Rev Cell Dev Biol, 2017, 33: 491-510. doi: 10.1146/annurev-cellbio-100616-060608 [55] Pribasnig M, Kien B, Pusch L, et al. Extended-resolution imaging of the interaction of lipid droplets and mitochondria [J]. Biochim Biophys Acta Mol Cell Biol Lipids, 2018, 1863(10): 1285-1296. [56] Gemmink A, Daemen S, Kuijpers H, et al. Super-resolution microscopy localizes perilipin 5 at lipid droplet-mitochondria interaction sites and at lipid droplets juxtaposing to perilipin 2 [J]. Biochim Biophys Acta Mol Cell Biol Lipids, 2018, 1863(11): 1423-1432. [57] Perkins H T, Allan V. Intertwined and finely balanced: Endoplasmic reticulum morphology, Dynamics, Function, and Diseases [J]. Cells, 2021, 10(9): 2341. doi: 10.3390/cells10092341 [58] Valm A M, Cohen S, Legant W R, et al. Applying systems-level spectral imaging and analysis to reveal the organelle interactome [J]. Nature, 2017, 546(7656): 162-167. doi: 10.1038/nature22369 [59] Reggiori F, Molinari M. ER-phagy: Mechanisms, regulation, and diseases connected to the lysosomal clearance of the endoplasmic reticulum [J]. Physiol Rev, 2022, 102(3): 1393-1448. doi: 10.1152/physrev.00038.2021 [60] Georgiades P, Allan V J, Wright G D, et al. The flexibility and dynamics of the tubules in the endoplasmic reticulum [J]. Sci Rep, 2017, 7(1): 16474. doi: 10.1038/s41598-017-16570-4 [61] Jung M, Mun J Y. Mitochondria and endoplasmic reticulum imaging by correlative light and volume electron microscopy [J]. J Vis Exp, 2019, 149: e59750. doi: 10.3791/59750 [62] Nixon-Abell J, Obara C J, Weigel A V, et al. Increased spatiotemporal resolution reveals highly dynamic dense tubular matrices in the peripheral ER [J]. Science, 2016, 354(6311): aaf3928. doi: 10.1126/science.aaf3928 [63] Schroeder L K, Barentine A, Merta H, et al. Dynamic nanoscale morphology of the ER surveyed by STED microscopy [J]. J Cell Biol, 2019, 218(1): 83-96. doi: 10.1083/jcb.201809107 [64] Guo Y, Li D, Zhang S, et al. Visualizing intracellular organelle and cytoskeletal interactions at nanoscale resolution on millisecond timescales [J]. Cell, 2018, 175(5): 1430-1442. doi: 10.1016/j.cell.2018.09.057 [65] Lewis S C, Uchiyama L F, Nunnari J. ER-mitochondria contacts couple mtDNA synthesis with mitochondrial division in human cells [J]. Science, 2016, 353(6296): f5549. doi: 10.1126/science.aaf5549 [66] Qiao C, Li D, Guo Y, et al. Evaluation and development of deep neural networks for image super-resolution in optical microscopy [J]. Nat Methods, 2021, 18(2): 194-202. doi: 10.1038/s41592-020-01048-5 [67] Gottschalk B, Klec C, Waldeck-Weiermair M, et al. Intracellular Ca2+ release decelerates mitochondrial cristae dynamics within the junctions to the endoplasmic reticulum [J]. Pflugers Arch, 2018, 470(8): 1193-1203. doi: 10.1007/s00424-018-2133-0 [68] Filipe A, Chernorudskiy A, Arbogast S, et al. Defective endoplasmic reticulum-mitochondria contacts and bioenergetics in SEPN1-related myopathy [J]. Cell Death Differ, 2021, 28(1): 123-138. doi: 10.1038/s41418-020-0587-z [69] Raiborg C, Wenzel E M, Pedersen N M, et al. Repeated ER-endosome contacts promote endosome translocation and neurite outgrowth [J]. Nature, 2015, 520(7546): 234-238. doi: 10.1038/nature14359 [70] Pavez M, Thompson A C, Arnott H J, et al. STIM1 Is required for remodeling of the endoplasmic reticulum and microtubule cytoskeleton in steering growth cones [J]. J Neurosci, 2019, 39(26): 5095-5114. doi: 10.1523/JNEUROSCI.2496-18.2019 -







 下载:
下载: