-
在传统的医学诊断与观察中,医学工作者将不同的“视觉色彩”赋予生物细胞组织样本从而区分细胞组织结构差异[1]。一方面,这些“色彩”由人工方式产生,并且都与某种特征结构对应,例如,化学染料染色[2]、荧光标记染色[3]等;另一方面,与自然环境中的色彩相比,这类医学成像领域的“视觉色彩”往往种类有限,经过多年的发展,产业规范不断迭代,产生方式基本已经被规范化和统一化,较为典型[4-9]。以组织学与病理学中使用最广泛的苏木精-伊红(H&E)染色法[10]为例:细胞核区域反映为蓝紫色外观,细胞基质和细胞质部分反映为粉红色外观[11-15]。这意味着颜色信息与其生物形态学特征密切相关。而荧光标记染色也具有类似特点。因此,正是因为结构与色彩之间的联系,才使得光学成像与计算成像的科研工作人员可以通过计算后处理的方式[16]降低生物医学成像处理样品的步骤,避免染色时的操作对最终结果产生影响[17],实现数字组织学图像[18]与传统医学金标准(Gold-Standard)——明场彩色显微成像[19]一致的彩色成像效果。
随着近些年人工智能技术的软/硬件的快速发展[20-24],基于人工智能色彩迁移技术的生物医学成像技术也得到了快速发展[25]。深度学习色彩迁移技术[26]指通过深度学习的方式,实现将目标图像色彩风格依照源图像色彩风格进行转换的色彩迁移技术。色彩迁移在生物医学成像方面有着较高的应用价值,如超声成像[27-28]、光声显微成像[29]、中远红外成像[30]等。文中将首先介绍几种深度学习色彩迁移的技术原理,列举此类技术在生物医学成像领域中的部分应用,最后展望人工智能色彩迁移在生物医学成像领域未来的发展方向。
-
色彩迁移[31]指两幅图像之间色彩的转换,通俗来说,就是一幅图像作为颜色的来源,称为源图像,另一幅图像作为待处理图像,称为目标图像,通过算法使目标图像在保留自身内容的情况下,拥有前者的色彩信息,即图像色彩风格的转换。
事实上,图像色彩迁移并不是一个新概念,在传统图像处理领域,已经有一些研究者在色彩迁移上取得了相对较好的效果,但其仍是处于对图像像素值处理的层面上,没有与图像内容加强联系,视觉上难以做到逼真的效果,如Reinhard等人[32] 提出了一组适用于各颜色分量的色彩迁移公式;Welsh等人[33]在Reinhard等人算法的基础上,利用查找匹配像素来实现灰度图像的色彩迁移。
而近些年,基于深度学习(Deep Learning, DL)的色彩迁移技术作为一种极具发展潜力的色彩迁移技术已经成为研究重点之一[34],然而色彩迁移仍被看作图像转换的范畴,其中心思想其实就是基于输入图像得到想要的输出图像的过程,是图像与图像之间的一种映射[35],那么如何实现此种映射,就成了研究者们关注的重点,同时,不同的切入点带来研究者们对此问题不同的理解,以及许多经典的算法和具体应用案例,如图像艺术风格的迁移[36]、灰度图上色[37]等。
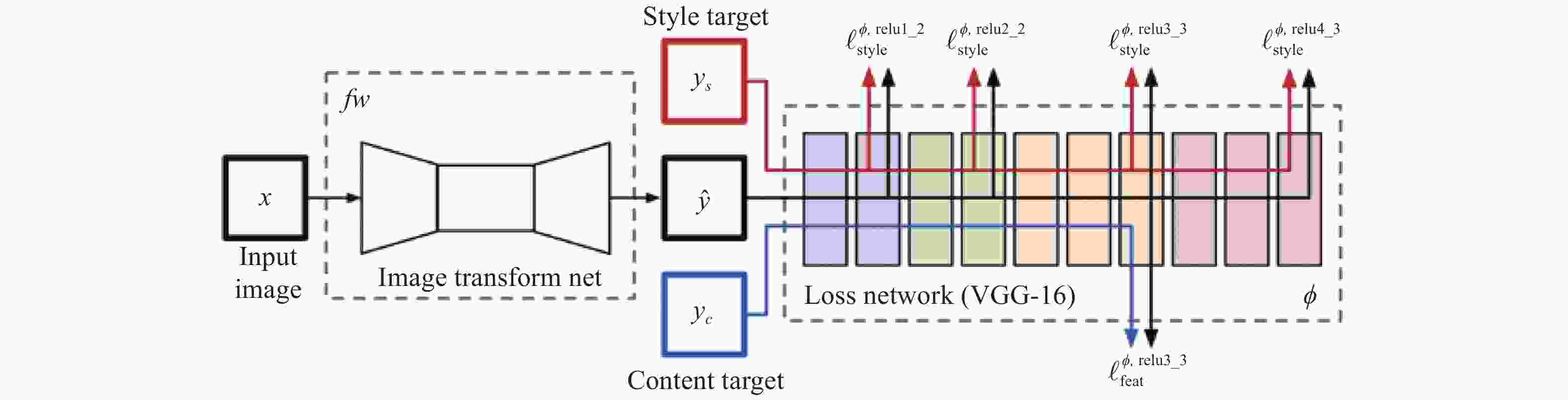
2015年,Gatys等人[38]受到卷积神经网络[39-44]的启发,首次将其应用于图像风格转换的问题上,提出了一种基于卷积神经网络(Convolution Neural Network, CNN)的图像风格迁移算法。该算法利用了卷积神经网络可以有效地提取到图像的内部特征[45],通过网络前馈传播[46]分层处理视觉信息并表达,在网络模型中抽象分离出了图像的内容信息与风格信息,进一步统计特征,迭代更新原始图像数据,直至风格信息误差收敛。值得一提的是算法在训练的过程中与常规CNN不同,该算法根据损失计算出的是输入图像的损失梯度,从而更新图像像素值,而不是常规CNN的模型参数。如图1所示。
算法的具体实施方式是把内容图像与风格图像以及白噪声输入至VGG网络中,则每层网络都会得到${N_l}$张${M_l}$大小的特征图,其个数取决于滤波器个数。在内容重建部分,将每一层中的特征图向量化后保存至一个矩阵${F^l} \in {\mathbb{R}^{{N_l} \times {M_l}}}$中,并希望使生成图像$\overrightarrow x $在该层的特征矩阵${p^l}$与内容图像在该层特征矩阵${F^l}$相同,从内容损失反向传播优化噪声图像$\overrightarrow x $,其内容损失如下:
$$ {L_{content}}(\overrightarrow p ,\overrightarrow x ,l) = \frac{1}{2}\sum\limits_{i,j} {{{\left( {F_{ij}^l - P_{ij}^l} \right)}^2}} $$ (1) 式中:$\overrightarrow p $为原始图像;$F_{ij}^l$为第$l$层$j$位置的第$i$个滤波器的响应。
在风格重构部分,作者使用了Gram矩阵统计图像的风格信息,包含图像的纹理与颜色特征,其中$ {G_{ij}}^l $是特征矢量$i$和$j$的内积:$ {G_{ij}}^l = \sum\nolimits_k {F_{ik}^l} F_{jk}^l $,因而对于风格图像$\overrightarrow a $与生成图像$\overrightarrow x $而言,由两者在第$l$层的Gram矩阵$ {A^l} $和$ {G^l} $即可定义该层的风格损失:
$$ {E_l} = \frac{1}{{4N_l^2M_l^2}}\sum\limits_{i,j} {{{({G_{ij}}^l - {A_{ij}}^l)}^2}} $$ (2) 进而得到总的风格损失:
$$ {L_{style}}(\overrightarrow a ,\overrightarrow x ) = \sum\limits_{l = 0}^L {{\omega _l}} {E_l} $$ (3) 式中:$ {\omega _l} $是style层的权重,取style层数的倒数,其他层权重为0。
对于风格迁移的部分来说,算法要在生成图像的过程中将内容损失与风格损失最小化,那么不难想到总的损失函数可定义为:
$$ {L_{total}}(\overrightarrow p ,\overrightarrow a ,\overrightarrow x ) = \alpha {L_{content}}(\overrightarrow p ,\overrightarrow x ,l) + \beta {L_{style}}(\overrightarrow a ,\overrightarrow x ) $$ (4) 式中:$\alpha $与$\;\beta $分别为内容与风格重构的权重因子。
Gatys等人的工作意义在于,其提示了可以使用卷积神经网络将图像特征抽象出来并做出处理,而不是手工建立一个数学或者统计模型,因而大大拓展了基于传统风格迁移研究的实际应用。尽管此算法在图像风格迁移工作中有着较为不错的效果,但显而易见地,其训练过程中优化的对象是噪声图像,存在大量迭代计算的步骤,因此也是极其耗时的,而事实上也是如此,该算法无法对单张图像实时迁移,以GTX2080 Ti为例,在使用单GPU加速的硬件条件下,对于内容复杂度不同的图像,其处理时间约为12~15 min。
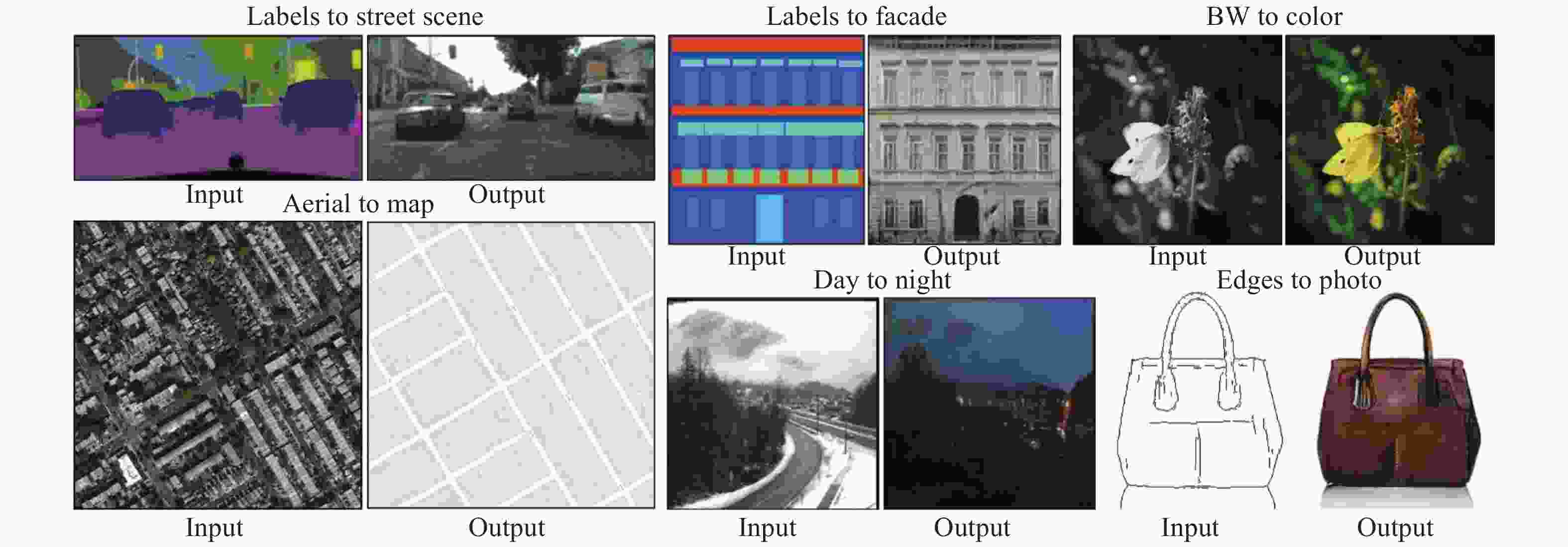
对于Gatys等人的风格迁移算法,其最大的缺点是在线迭代时间过长,导致算法难以得到更为广泛的实际生产应用。而Johnson等人[47]在2016年发表的成果弥补了这一缺点,其核心思想是规避大量的在线迭代运算,通过使用感知损失函数,直接训练出一个“端到端”的网络模型,在测试阶段通过将内容图片作为网络的输入得到风格化的结果图片,由于该算法在生成阶段只需要进行一次前向传播且不需要计算梯度用以更新网络权重或初始化图片,与基于图片优化的方法相比,在生成效果不变的同时,生成速度大约提升了1000倍。然而,这样的预训练网络最初是为物体识别而设计的,因此深层特征往往专注于主要目标而忽略其他细节,生成的图像通常不令人满意[48]。如图2所示。
另一方面,生成对抗网络[49](GAN)在图像生成领域也展现出强大的能力[50-56]。GAN的基本思想是一种二人零和博弈思想。网络通常需要构建生成模型(G)与判别模型(D),生成器用来对输入的样本\噪声做处理,将它生成为一个逼真的样本;判别器则作为一个有监督的二分类器,对真实样本与生成样本做出区分。在训练的过程中,生成器尽可能地将输出逼近真实样本的分布,而判别器将尽可能地区分真实样本与生成样本,生成器与判别器二者对抗博弈,其优化过程可看作为一个最大最小化问题:
$$\begin{split} \mathop {\min }\limits_G \mathop {\max }\limits_D V(D,G) =& {E_{x \sim Pdata(x)}}[\log (D(x))] +\\ &{E_{z\sim Pz(z)}}[\log (1 - D(G(z)))] \end{split} $$ (5) 正由于GAN表现出可自动学习目标样本集的真实样本分布的能力,一些基于GAN的图像转换算法相继被提出并取得了很好的效果。2017年,pix2 pix算法[57]被提出发表在CVPR2017(图3)。该算法基于条件GAN(cGAN)[58],相比于传统GAN,pix2 pix不再使用噪声作为输入图像,而使用用户输入的图像进行映射,成功地实现了色彩风格的迁移转换,这种对应关系在训练过程的拟合则需要成对的图像做训练数据,通过生成器与鉴别器的对抗博弈,以最大化生成器的能力与最小化鉴别器的能力作为学习目标来获得这种对应关系。显而易见地,由于得到这种有固定特征关系的映射,网络的应用范围是比较广泛的,如基于特定目标的图像转换、色彩迁移等。
尽管pix2 pix已经具备了获取图像之间映射转换关系的能力,但由于其训练样本需要成对的图像,而现实生活中成对的数据是很难获得的,因此极大地限制了pix2 pix的应用。随后,pixp2 pix的研究者们为解决该问题,提出了一种更加强大的网络——循环一致生成对抗网络(CycleGAN)[59]。CycleGAN不需要成对的图像作为网络的训练样本,只需要将训练样本、目标样本的集合输入至网络中,通过训练拟合,即可获得两个集合之间的映射生成关系。如图4所示。
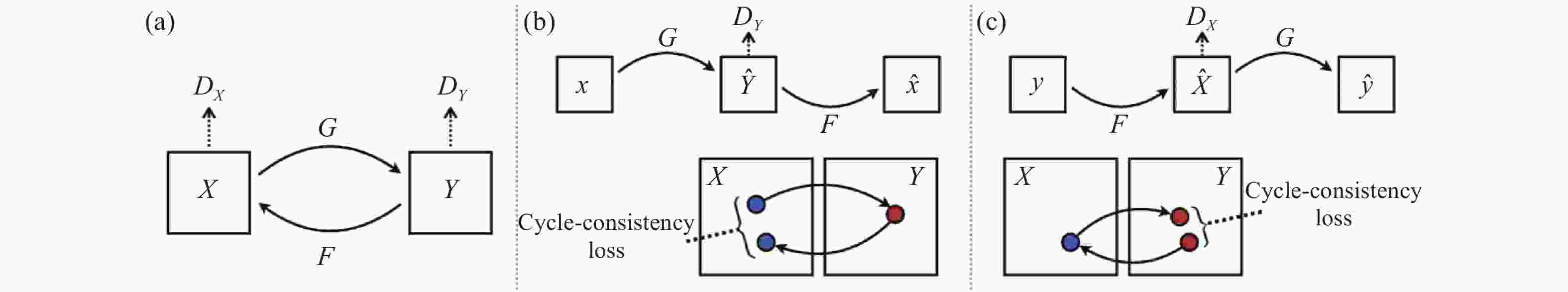
虽然CycleGAN摒弃了pix2 pix的成对训练数据,但同时,CycleGAN也就因此放弃了网络对输出的限制条件,这无疑会使网络的输出图像呈现随机性,因此,作者在构建网络时除经典GAN网络的对抗损失外,为网络添加了循环一致损失,来保证生成的图像保留原始图像的特性限制[60]。其网络结构如图5所示。
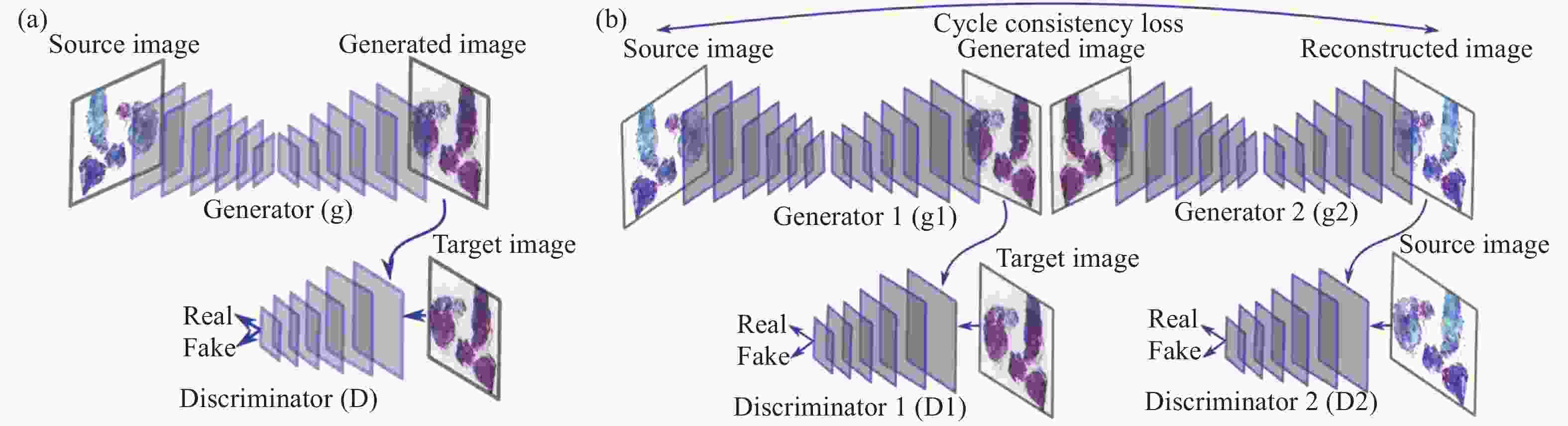
图中,G与F是生成器,${D_X}$与${D_Y}$是鉴别器,其循环一致损失通过限制X域到Y域再由Y域到X域的映射有效地保证了在图像生成过程中不会丢失原始图像的语义信息。 与此同时,CycleGAN不易改变图像内容形状的特点也非常适合应用于色彩迁移问题上。如图6所示。
-
在生物医学的临床实践中,组织或细胞的细胞学切片的显微评估一直是病理学诊断中最为重要的一部分[61],如涂片、手术过程中肿瘤组织边界分析和死后的组织学检查,但是随着医疗需求的日益增长,传统检测观察方式已经难以满足当今的需要[62]。然而,近些年人工智能深度学习领域的发展促成了计算机辅助分析领域与临床医疗的有效结合,其在生物医学成像分析上也逐渐表现出较高的发展潜力[63]。
-
在病理学的研究中,医务工作者们往往通过使用化学染液的浸染方式,将不同的颜色赋予生物组织切片的不同结构来区分组织,这些颜色数量有限[64],且与其生物结构存在直接联系。这些“色彩——结构”的简单映射就提示了研究者们可以将结构特征与颜色特征相联系[65],由计算机构建出由结构到色彩的映射模型,从而对未经化学染色的生物切片的图像虚拟染色,以降低成本,提高观察效率。
Pranita等人[66]提到了在有监督和无监督方法中使用深度学习模型,用非线性多模式(NLM)成像方法观察冷冻切片并进行计算染色,将不同模式下的图像自动转换为H&E染色图像,使观察标本不再局限于石蜡包埋的FFPE染色切片。监督学习方法使用成对图像的pix2 pix模型,无监督学习方法使用循环CGAN模型。对于前者,需要一对对应的NLM图像和组织病理学染色的H&E图像。因此,组织病理学染色的H&E图像的图像配准至关重要。另一方面,循环CGAN模型不需要NLM图像和组织病理学染色的H&E图像一一对应。因此,减少了图像配准和病理染色的工作量。如图7所示。
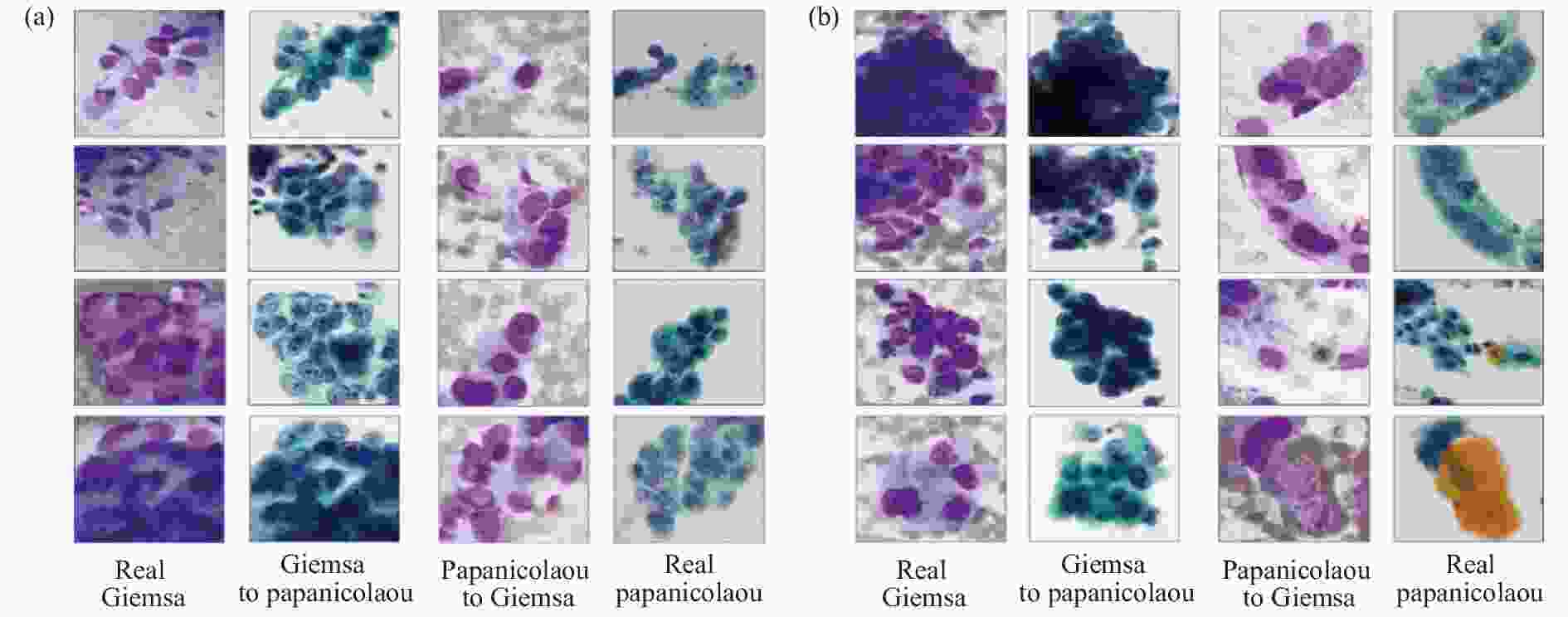
然而监督学习方法在实际应用中会遇到训练样本的制约,如研究者们很难获得对同一观察样本片进行多种标准染色处理的成对图像,以供网络训练。而无监督的学习方法在训练数据的获得上更具优势。其中,CycleGAN这一无监督学习[67]模型,自诞生以来便表现出了其从源域图像到目标域图像强大的转化能力、出色的纹理结构保留能力,且无需成对的训练样本,这使它成为了一种生物医学成像领域虚拟染色的流行方法。Teramoto等人[68]使用 CycleGAN 在细胞学图像中进行了巴氏染色[69]和吉姆萨[70]染色之间的相互染色转换,实验效果如图8所示。经过视觉评估,虽然实际图像与转换后的图像相比表现出更好的观察效果,但算法结果仍具有潜在的价值,如可以处理不同的染色剂不能应用于同一个细胞的问题等。
传统病理学检测中,涂片、染色、镜检等过程限制着病情的诊断速度[71],且操作的规范性与实验室观察条件也影响着医生判断的准确性[72]。尽管基于深度学习的色彩迁移技术体现出较为高效的处理能力与良好的染色效果,但最终检测效果仍一定程度上依赖于下游工作者对染色的生物样本进行显微观察分析,从而给出病理诊断。这一过程中,医务人员的知识技能与经验技术就显得尤为重要,并且影响着最终病理诊断。对于这个问题,基于深度学习色彩迁移的切片色彩标准化技术与深度学习识别网络的结合,在检测速度与准确率上体现出了较大的优势。Lo等人[73]使用CycleGAN对肾小球进行H&E染色,结果输入至faster R-CNN中检测肾小球,并由四名医生评估(如图9所示)。实验结果表明,医生无法区分真实染色和迁移染色,自动肾小球检测方法优于医生手动标记的方法。该方法效果良好,提高了肾脏病理诊断的效率。这项工作有助于自动化医疗诊断领域。

图 9 Lo等人的工作[73]。(a) Lo等人的CycleGAN结构;(b) Lo等人的faster R-CNN结构;(c) 使用不同的 H&E 训练模型来测试不同点的图像得到的P-R曲线,其中“O”和“×”分别表示由四名医生对H&E和PAS图像进行人工检测的结果
Figure 9. Work done by Lo et al[73]. (a) CycleGAN structure of Lo et al; (b) The faster R-CNN structure of Lo et al;(c) P–R curves using different H&E trained models to test images with different stains, where “O” and “×” denote the manual detection results of H&E and PAS images, respectively, performed by four doctors
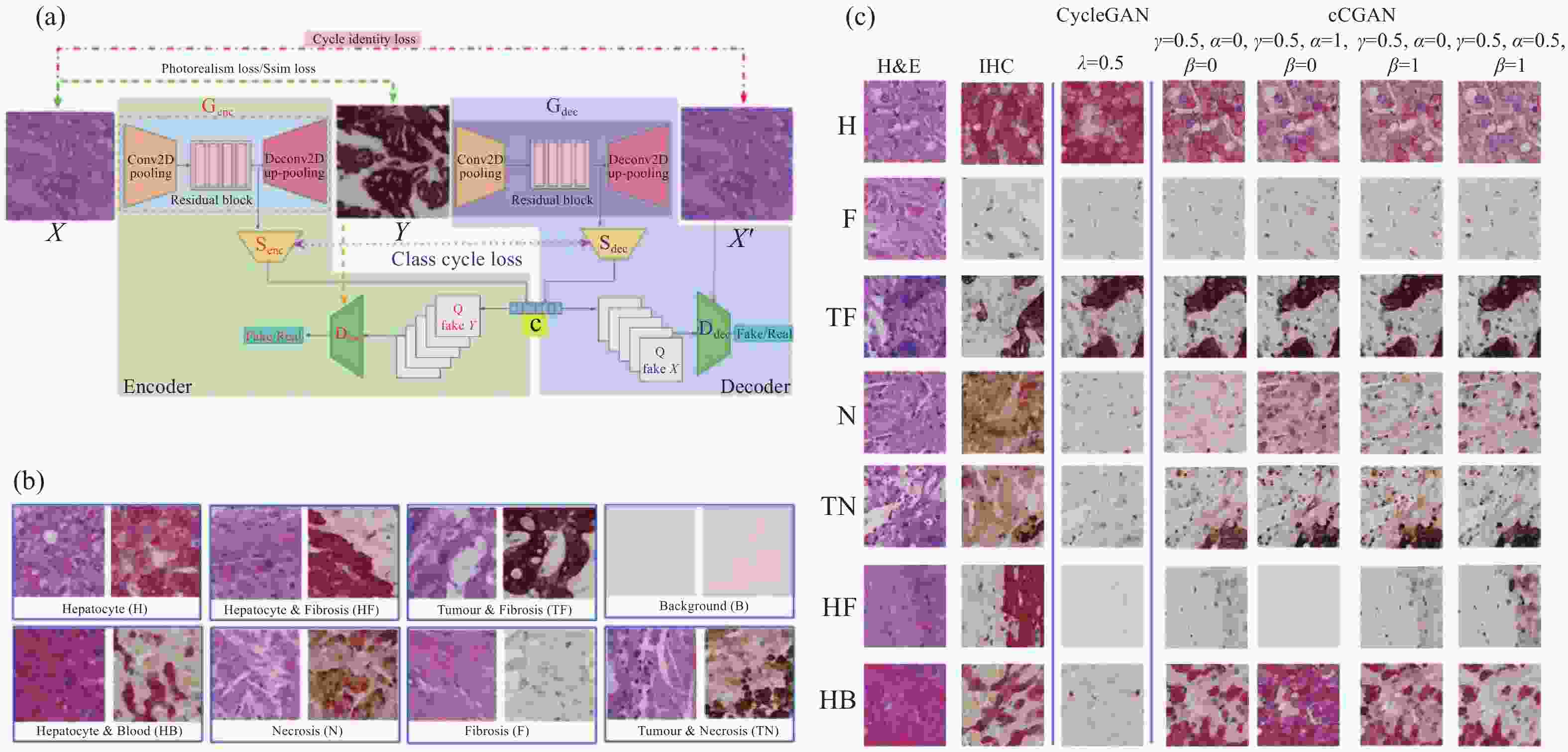
在不同的医学背景下,分析单一染色模式的切片难以对患者的病情给出较为全面的评估。不同的应用场景下,研究人员往往需要针对不同的染色模式对网络模型做出改进与取舍。例如,相比H&E染色图像,IHC染色图像具有更高的对比度[74],Xu等人[75]提出了名为cCGAN的网络模型,对CycleGAN进行了改进,通过添加类别条件和引入两个结构损失函数,成功将H&E染色图像转换为IHC染色图像,实现了多子域翻译并提高翻译准确性,便于在同一载玻片上进行虚拟IHC染色,降低了检测成本(图10)。de Bel等人[76]对CycleGAN进行了简单而有效的更改,将生成器网络的任务转向学习差异映射,即源域和目标域的残差。这允许生成器网络仅专注于域适应,同时保持形态完整性作为参考,其结构如图11所示。
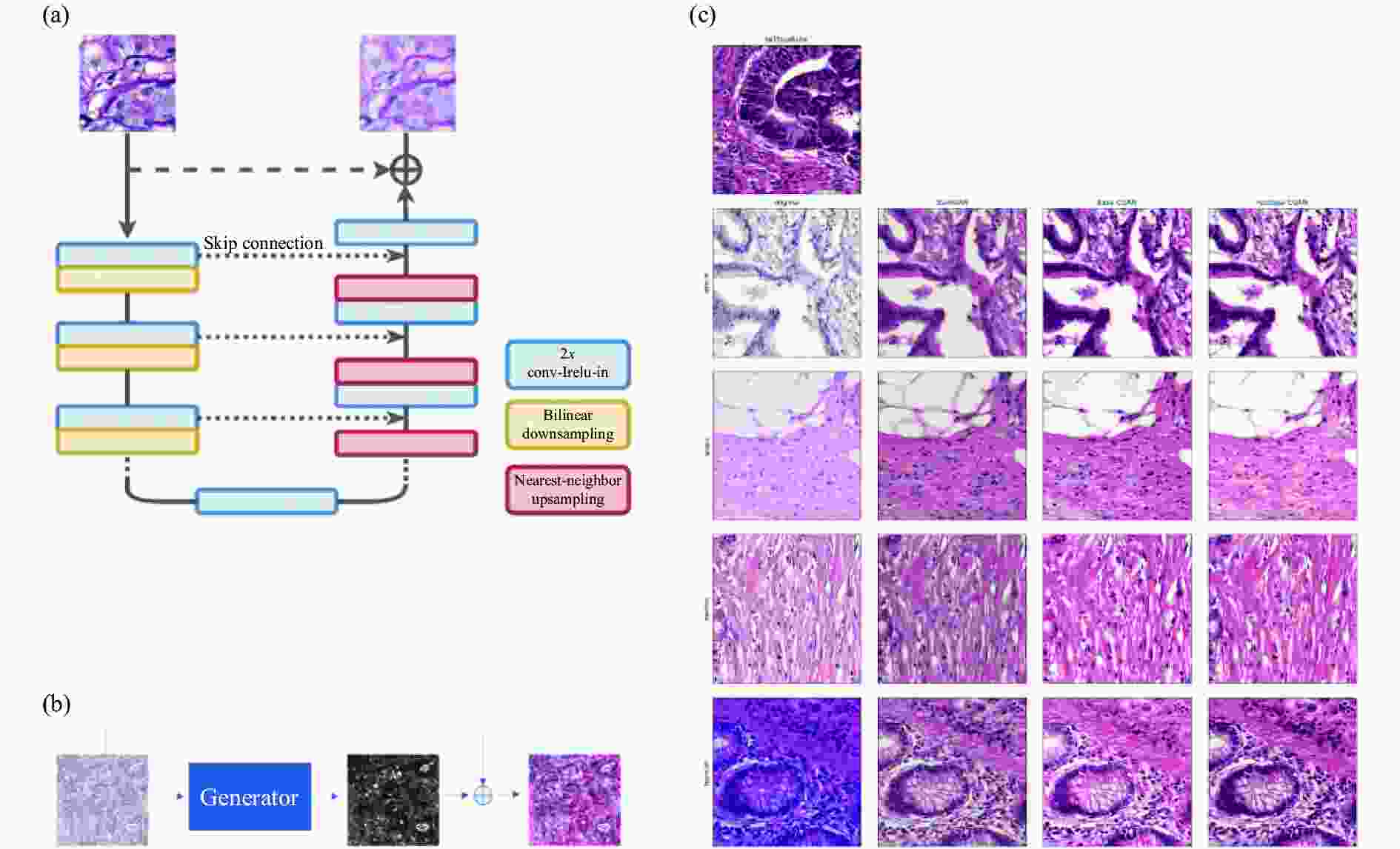
图 11 de Bel等人的工作[76]。(a) 残差CycleGAN中生成器的架构,与标准U-net非常相似;(b) 生成器学习源域和目标域之间的差异映射或残差;(c) 使用CycleGAN方法转化前后的结肠组织样本
Figure 11. Work done by de Bel et al[76]. (a) Architecture of the generator in the residual CycleGAN, closely resembling the standard U-net; (b) The generator learns the difference mapping or residual between a source and target domain; (c) Samples of colon tissue before and after transformation with the CycleGAN approaches
基于深度学习的色彩迁移技术不仅在图像后处理上取得良好的效果,其也可结合不同的医学图像信息获取技术实现更好的应用效果,从而扩展技术应用场景。对于传统的组织病理学来说,组织学图像的获取往往需要复杂的处理步骤,其中过程包括切片、标本固定、以及组织学染色等[77-79],过程会花费大量时间,且切片厚度影响着后续的染色与观察结果。Kang等人[80]通过结合紫外光声显微镜[81](UV-PAM)与CycleGAN,提供了一种快速且无标记的组织学成像方法Deep-PAM,不仅可以获得薄脑切片的虚拟 H&E 染色图像,也能够获得厚的和新鲜的无标签标本表面的虚拟H&E染色图像。这种Deep-PAM成像和虚拟染色方法可用于传统组织学以替代组织化学H&E染色过程,从而节省染色时间,同时,由于新鲜组织是直接成像,不需要组织处理和化学染色,节省了大量的成本,如组织处理机器、人工和试剂支出。如图12所示。
尽管基于深度学习的色彩迁移方法有着许多传统方法难以比拟的实际效果[82],但是仍存在模型泛化能力不足的问题[83-85],在充分考虑到这一点后,Chen等人[86]提出了一种无监督的方法来规范细胞病理学图像样式,其结构如图13所示。
该网络是由一个样式删除模块和一个样式重建模块构建,前者将源图像和目标图像具有大致一致的分布,而后者将分布重建到源域,通过颜色编码掩码和 L1 损失来强制色调和结构的一致性,并通过域对抗风格重建确保来自不同域的归一化图像的风格与用户选择的风格一致。文章中对比了几种经典归一化方法(SPCN[87]、Macenko[88]、Reinhard[32]、Khan[89]、Gupta[90]、Zheng[91]、CycleGAN[59]、StainGAN[92] 和Tellez[93])的实验效果,结果对比如图14所示。
-
基于深度学习的色彩迁移生物医学成像技术在对数字图像信息的处理上体现出良好的效果,在与光学成像结合的应用过程中也展现出强大的生命力。其为研究者提供较为良好的观察效果,同时,也在一定程度上降低了硬件要求。
在显微成像领域,目前没有成像镜头的无透镜全息显微镜可以用较为低的成本实现大视场成像。PIE(Ptychographic Iterative Engine)迭代重建算法是一种衍射成像方法,它可以通过移动光源或生物医学病理样本载玻片来扩展视场(FOV),开发低成本的大视场无透镜显微镜。在PIE中,照明光要求具有高/部分相干性,因此,要获得彩色显微镜图像,在传统的PIE中需要具有不同主波长值的3个以上照明光源。笔者所在课题组提出了一种基于计算深度学习色彩迁移方法的改进PIE,以实现彩色大FOV无透镜显微镜成像[94]。该方法中,仅使用一个高/部分相干光源进行照明,其中图像数据比彩色PIE显微镜在多重照明下的图像少3倍。该网络是在H&E染色的病理组织切片的基础上进行训练的。因此,它仅适用于H&E染色的病理组织载玻片。但深度学习色彩迁移方法同样适用于其他染色医学诊断,例如荧光染色、苏丹染色等。如图15所示。
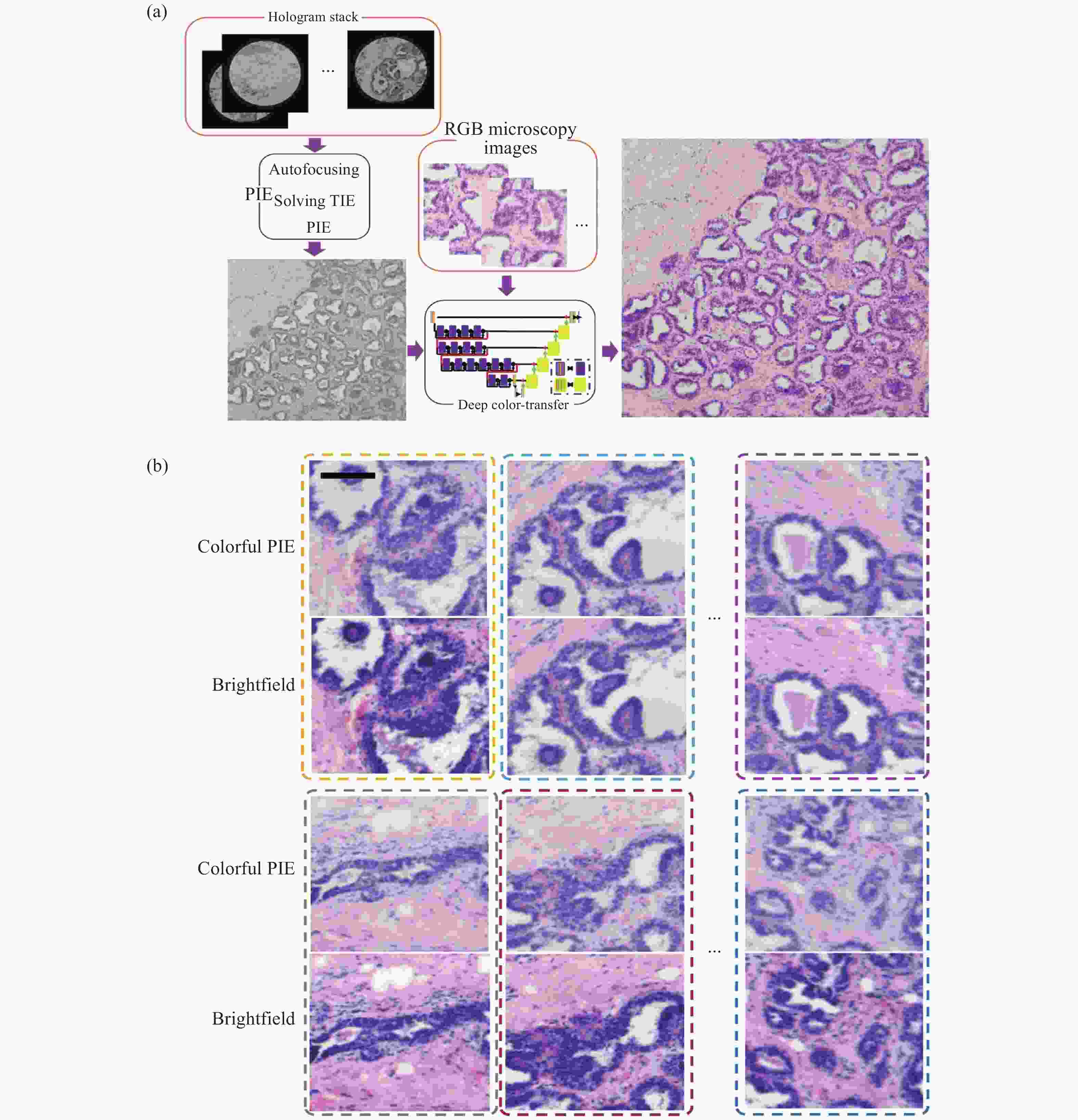
图 15 深度学习彩色PIE无透镜衍射显微镜[94]。(a) 仅单色照明的彩色PIE显微镜计算算法流程图;(b) 彩色PIE显微图像和传统RGB明场图像比较
Figure 15. Deep learning colorful PIE lens-less diffraction microscopy[94]. (a) Flow charts of computational algorithms for colorful PIE microcopy with only one kind illumination; (b) Vision comparisons of colorful PIE microscopy images and conventional RGB brightfield images
传统的无透镜全息显微镜为了获得彩色图像,应使用至少3个离散波长的准色光源,如红色 LED (R)、绿色 LED (G) 和蓝色 LED (B)。笔者所在课题组通过深度学习实现虚拟染色[95],将灰色无透镜显微镜图像转换为彩色图像(图16)。通过输入绿色 LED@550 nm 照明下的灰度无透镜显微镜图像和彩色明场显微镜图像,训练生成对抗网络 (GAN),并将其应用于H&E染色病理组织样本成像。该方法运行稳定,并且无需任何数据和硬件成本即可将无灰度透镜的片上显微镜图像扩展为彩色的人类视觉图像,可以用于改进无透镜显微镜在远程病理学和资源有限情况下的应用。

图 16 虚拟染色的无透镜片上显微镜[95]。(a) 无透镜片上显微镜;(b) 实现虚拟彩色无透镜片上显微镜的数据处理,黄色比例尺为200 μm;(c) 建立深度学习GAN网络以实现虚拟着色;(d) 无透镜片上显微图像、台式商用显微图像和虚拟着色图像的比较
Figure 16. Virtual colorful lens-free on-chip microscopy[95]. (a) Lens-free on-chip microscope; (b) Data process to achieve virtual colorful lens-free on-chip microscopy. The yellow scale bar is 200 μm; (c) Deep learning GAN network established to achieve virtual colorization; (d) Comparisons of lens-free on-chip microscopy image, bench-top commercial microscopy image and virtual colorization image
另一方面,在光学成像系统中,单透镜无需精密组装、校准和测试,有利于便携式和低成本显微镜的发展,然而,由一种材料制成的单透镜难以平衡光谱色散或色差。为解决这个问题,笔者课题组结合单透镜显微方法和计算成像提出了一种基于深度学习图像风格转移算法的新方法[96],并用于解决临床病理切片显微镜中的这一问题(图17)。该方法使用了具有高截止频率和线性信号特性的单透镜非球面透镜,通过训练好的深度学习网络增强可以很容易地将单色灰度显微图片转换为彩色显微图片,单色CMOS图像传感器只记录一次。实验证明了对于H&E染色的病理组织切片,绿色LED照明和红色LED照明将提供比蓝色LED照明更好的 PSNR和SSIM。此外,虚拟着色方法也适用于其他单透镜显微方法和其他色差成像系统,例如传统的球面透镜、衍射透镜和超表面透镜。可以相信,单透镜显微镜的计算虚拟着色方法将推动低成本、便携式单透镜显微方法在医学病理标记染色观察(如H&E染色、荧光标记等)生物医学研究中的发展。
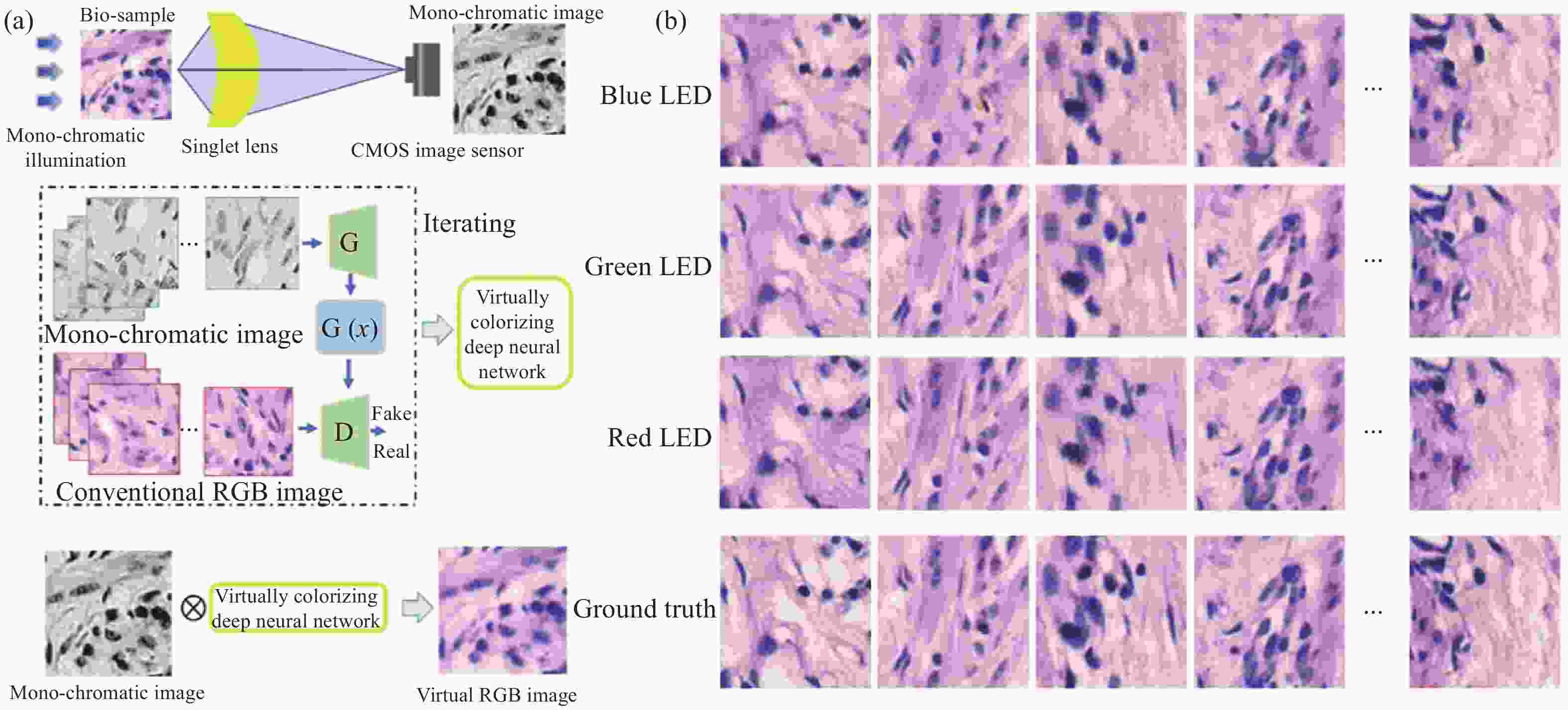
图 17 单线态显微镜着色[96]。(a) 实现单线态显微镜着色的概述;(b) B/G/R照明下的200组图像,以评估虚拟染色显微镜图像的平均PNSR和SSIM
Figure 17. Singlet microscopy colorization[96]. (a) An overview to achieve the singlet microscopy colorization; (b) 200 group images under B/G/R illumination to evaluate the virtual colorized microscopy images’ average PNSRs and SSIMs
-
过去十年,人工智能得到了快速发展,并应用于各个学科的研究领域当中,基于人工智能色彩迁移的生物医学成像技术是计算机辅助分析与生物成像领域交叉融合的结果,它给传统的生物医学成像领域提供了一个新的发展方向,是一种极具前景的应用技术。文中综述了近年来几种深度学习色彩迁移的技术原理,列举了此类技术在生物医学成像领域中的部分应用,如对组织病理学切片数字图像的色彩迁移、无透镜和单透镜成像的虚拟彩色增强等。按照应用领域、网络结构、学习方法和所解决问题的不同,表1给出了色彩迁移技术的使用情况统计结果。由表1可见,CycleGAN结构和无监督学习类型的深度学习网络在解决各种问题的应用场景中占了较大的比重。如前所述,CycleGAN所表现出的强大性能使其在生物医学成像虚拟染色领域得到广泛应用。其从源域图像到目标域图像强大的转化能力、出色的纹理结构保留能力,使得虚拟染色图像可以有效地应用于组织病理学分析当中。而无监督学习的特点使得研究者无需提供成对的训练样本,减少了研究人员构建符合要求的训练样本数据库的工作,从而大大降低了算法的使用门槛,因此相比监督学习的网络,它拥有着更加广泛的应用。
表 1 色彩迁移技术的使用情况统计
Table 1. Statistics on the usage of color transfer technology
Application field Network structure Learning method Application problems Supervised learning Unsupervised learning Color transfer technology of pathological section images pix2 pix √ Computational tissue staining Cycle CGAN √ Computational tissue staining CycleGAN √ Mutual stain CycleGAN,
Faster R-CNN√ √ Tissue staining and detection cCGAN,
Residual CycleGAN√ Model improvements for different demand backgrounds Deep-PAM √ Combining different medical image information acquisition technologies GAN √ Unsupervised image style normalization Virtual color enhancement for lensless and single lens imaging GAN √ PIE GAN √ Improvements in lensless microscopes U-Net √ Computational virtual shading method for single lens microscopy 尽管这些新的技术方法在处理生物医学成像问题中取得了令人满意的效果,但其在具体应用中尚有问题亟待解决。首先,基于深度学习的色彩迁移网络面临着模型泛化能力较差的问题,网络的泛化能力又极大地依赖于训练数据集的内容丰富性。作为一个“端到端”的复杂非线性映射关系,其结果输出对输入图像的质量要求较高,在不同照明环境下,网络的输出结果很难保持较高的稳定性。而这类应用要求在医学工作中难以完全满足,如切片在不同显微设备间切换观察、切片制作时的薄厚差异等带来的照明差异。这也限制了算法模型被安全有效地应用于临床工作中。另一方面,对于医学图像的色彩迁移结果评价仍未建立合理有效的评判标准,这在一定程度上限制着这类新技术在具体应用中的发展推进。因此,构建更加完备的训练样本库、结合新的信息处理技术、建立起统一的结果评价标准等,将是未来打破限制、发展延伸此类技术的重点。
Deep learning-based color transfer biomedical imaging technology
-
摘要: 传统病理学检测中,由于复杂的染色流程和单一的观察形式等限制着病情的诊断速度,而染色过程实质上是将颜色信息与形态特征关联,效果等同于现代数字技术的生物医学图像的图义分割,这使得研究者们可以通过计算后处理的方式,大大降低生物医学成像处理样品的步骤,实现与传统医学染色金标准一致的成像效果。近些年人工智能深度学习领域的发展促成了计算机辅助分析领域与临床医疗的有效结合,人工智能色彩迁移技术在生物医学成像分析上也逐渐表现出较高的发展潜力。文中回顾了深度学习色彩迁移的技术原理,列举此类技术在生物医学成像领域中的部分应用,并展望了人工智能色彩迁移在生物医学成像领域的研究现状和可能的发展趋势。Abstract: In traditional pathology detection, the speed of diagnosis is limited due to the complex staining process and single observation form. The staining process is essentially associating color information with morphological features, and the effect is equivalent to that of biomedical images of modern digital technology. Sense segmentation, which allows researchers to greatly reduce the steps of biomedical imaging processing samples through computational post-processing, and achieve imaging results consistent with the gold standard of traditional medical staining. In recent years, the development of artificial intelligence deep learning has contributed to the effective combination of computer-aided analysis and clinical medicine, and artificial intelligence color transfer technology has gradually shown high development potential in biomedical imaging analysis. This paper will review the technical principles of deep learning color transfer, enumerate some applications of such technologies in the field of biomedical imaging, and look forward to the research status and possible development trends of artificial intelligence color transfer in the field of biomedical imaging.
-
Key words:
- deep learning /
- artificial intelligence /
- color transfer /
- biomedical imaging
-
图 9 Lo等人的工作[73]。(a) Lo等人的CycleGAN结构;(b) Lo等人的faster R-CNN结构;(c) 使用不同的 H&E 训练模型来测试不同点的图像得到的P-R曲线,其中“O”和“×”分别表示由四名医生对H&E和PAS图像进行人工检测的结果
Figure 9. Work done by Lo et al[73]. (a) CycleGAN structure of Lo et al; (b) The faster R-CNN structure of Lo et al;(c) P–R curves using different H&E trained models to test images with different stains, where “O” and “×” denote the manual detection results of H&E and PAS images, respectively, performed by four doctors
图 11 de Bel等人的工作[76]。(a) 残差CycleGAN中生成器的架构,与标准U-net非常相似;(b) 生成器学习源域和目标域之间的差异映射或残差;(c) 使用CycleGAN方法转化前后的结肠组织样本
Figure 11. Work done by de Bel et al[76]. (a) Architecture of the generator in the residual CycleGAN, closely resembling the standard U-net; (b) The generator learns the difference mapping or residual between a source and target domain; (c) Samples of colon tissue before and after transformation with the CycleGAN approaches
图 15 深度学习彩色PIE无透镜衍射显微镜[94]。(a) 仅单色照明的彩色PIE显微镜计算算法流程图;(b) 彩色PIE显微图像和传统RGB明场图像比较
Figure 15. Deep learning colorful PIE lens-less diffraction microscopy[94]. (a) Flow charts of computational algorithms for colorful PIE microcopy with only one kind illumination; (b) Vision comparisons of colorful PIE microscopy images and conventional RGB brightfield images
图 16 虚拟染色的无透镜片上显微镜[95]。(a) 无透镜片上显微镜;(b) 实现虚拟彩色无透镜片上显微镜的数据处理,黄色比例尺为200 μm;(c) 建立深度学习GAN网络以实现虚拟着色;(d) 无透镜片上显微图像、台式商用显微图像和虚拟着色图像的比较
Figure 16. Virtual colorful lens-free on-chip microscopy[95]. (a) Lens-free on-chip microscope; (b) Data process to achieve virtual colorful lens-free on-chip microscopy. The yellow scale bar is 200 μm; (c) Deep learning GAN network established to achieve virtual colorization; (d) Comparisons of lens-free on-chip microscopy image, bench-top commercial microscopy image and virtual colorization image
图 17 单线态显微镜着色[96]。(a) 实现单线态显微镜着色的概述;(b) B/G/R照明下的200组图像,以评估虚拟染色显微镜图像的平均PNSR和SSIM
Figure 17. Singlet microscopy colorization[96]. (a) An overview to achieve the singlet microscopy colorization; (b) 200 group images under B/G/R illumination to evaluate the virtual colorized microscopy images’ average PNSRs and SSIMs
表 1 色彩迁移技术的使用情况统计
Table 1. Statistics on the usage of color transfer technology
Application field Network structure Learning method Application problems Supervised learning Unsupervised learning Color transfer technology of pathological section images pix2 pix √ Computational tissue staining Cycle CGAN √ Computational tissue staining CycleGAN √ Mutual stain CycleGAN,
Faster R-CNN√ √ Tissue staining and detection cCGAN,
Residual CycleGAN√ Model improvements for different demand backgrounds Deep-PAM √ Combining different medical image information acquisition technologies GAN √ Unsupervised image style normalization Virtual color enhancement for lensless and single lens imaging GAN √ PIE GAN √ Improvements in lensless microscopes U-Net √ Computational virtual shading method for single lens microscopy -
[1] Chantziantoniou N, Donnelly A D, Mukherjee M, et al. Inception and development of the papanicolaou stain method [J]. Acta Cytologica, 2017, 61(4-5): 266-280. doi: 10.1159/000457827 [2] Fischer A H, Jacobson K A, Rose J, et al. Hematoxylin and eosin staining of tissue and cell sections [J]. Cold Spring Harbor Protocols, 2008, 2008(5): pdb.prot4986. doi: 10.1101/pdb.prot4986 [3] Beutner E H. Immunofluorescent staining: The fluorescent antibody method [J]. Bacteriological Reviews, 1961, 25(1): 49-76. doi: 10.1128/br.25.1.49-76.1961 [4] Irshad H, Veillard A, Roux L, et al. Methods for nuclei detection, segmentation, and classification in digital histopathology: A review—current status and future potential [J]. IEEE Reviews in Biomedical Engineering, 2013, 7: 97-114. [5] Chari S T, Echelmeyer S. Can histopathology be the “Gold Standard” for diagnosing autoimmune pancreatitis? [J]. Gastro-enterology, 2005, 129(6): 2118-2120. doi: 10.1053/j.gastro.2005.10.034 [6] Onder D, Zengin S, Sarioglu S. A review on color normalization and color deconvolution methods in histopathology [J]. Applied Immunohistochemistry & Molecular Morphology, 2014, 22(10): 713-719. [7] De Matos J, Britto Jr A S, Oliveira L E S, et al. Histopathologic image processing: A review [J]. arXiv preprint, 2019: 1904.07900. [8] Ursache R, Andersen T G, Marhavý P, et al. A protocol for combining fluorescent proteins with histological stains for diverse cell wall components [J]. The Plant Journal, 2018, 93(2): 399-412. [9] Taqi S A, Sami S A, Sami L B, et al. A review of artifacts in histopathology [J]. Journal of Oral and Maxillofacial Pathology: JOMFP, 2018, 22(2): 279. [10] Celis R, Romero E. Unsupervised color normalisation for H and E stained histopathology image analysis[C]//11th International Symposium on Medical Information Processing and Analysis. International Society for Optics and Photonics, 2015, 9681: 968104. [11] Abraham T, Shaw A, O'Connor D, et al. Slide-free MUSE microscopy to H&E histology modality conversion via unpaired image-to-image translation GAN models [J]. arXiv preprint, 2020: 2008.08579. [12] de Haan K, Rivenson Y, Wu Y, et al. Deep-learning-based image reconstruction and enhancement in optical microscopy [J]. Proceedings of the IEEE, 2019, 108(1): 30-50. [13] Liang H, Plataniotis K N, Li X. Stain style transfer of histopathology images via structure-preserved generative learning[C]//International Workshop on Machine Learning for Medical Image Reconstruction. Cham: Springer, 2020: 153-162. [14] Mahapatra D, Bozorgtabar B, Thiran J P, et al. Structure preserving stain normalization of histopathology images using self supervised semantic guidance[C]//International Conference on Medical Image Computing and Computer-Assisted Intervention. Cham: Springer, 2020: 309-319. [15] Shi M, McMillan K L, Wu J, et al. Cisplatin nephrotoxicity as a model of chronic kidney disease [J]. Laboratory Investigation, 2018, 98(8): 1105-1121. [16] Rivenson Y, Göröcs Z, Günaydin H, et al. Deep learning microscopy [J]. Optica, 2017, 4(11): 1437-1443. doi: 10.1364/OPTICA.4.001437 [17] Bilgin C C, Rittscher J, Filkins R, et al. Digitally adjusting chromogenic dye proportions in brightfield microscopy images [J]. Journal of Microscopy, 2012, 245(3): 319-330. doi: 10.1111/j.1365-2818.2011.03579.x [18] Veta M, Pluim J P W, Van Diest P J, et al. Breast cancer histopathology image analysis: A review [J]. IEEE Transactions on Biomedical Engineering, 2014, 61(5): 1400-1411. doi: 10.1109/TBME.2014.2303852 [19] Hooja S, Pal N, Malhotra B, et al. Comparison of Ziehl Neelsen & Auramine O staining methods on direct and concentrated smears in clinical specimens [J]. The Indian Journal of Tuberculosis, 2011, 58(2): 72-76. [20] Mak K K, Pichika M R. Artificial intelligence in drug development: present status and future prospects [J]. Drug Discovery Today, 2019, 24(3): 773-780. doi: 10.1016/j.drudis.2018.11.014 [21] Gunčar G, Kukar M, Notar M, et al. An application of machine learning to haematological diagnosis [J]. Scientific Reports, 2018, 8(1): 1-12. [22] Chen H, Engkvist O, Wang Y, et al. The rise of deep learning in drug discovery [J]. Drug Discovery Today, 2018, 23(6): 1241-1250. doi: 10.1016/j.drudis.2018.01.039 [23] Krittanawong C. The rise of artificial intelligence and the uncertain future for physicians [J]. European Journal of Internal Medicine, 2018, 48: e13-e14. doi: 10.1016/j.ejim.2017.06.017 [24] Grys B T, Lo D S, Sahin N, et al. Machine learning and computer vision approaches for phenotypic profiling [J]. Journal of Cell Biology, 2017, 216(1): 65-71. doi: 10.1083/jcb.201610026 [25] Rivenson Y, Liu T, Wei Z, et al. PhaseStain: The digital staining of label-free quantitative phase microscopy images using deep learning [J]. Light: Science & Applications, 2019, 8(1): 1-11. doi: 10.1038/s41377-019-0129-y [26] Shaban M T, Baur C, Navab N, et al. Staingan: Stain style transfer for digital histological images[C]//2019 IEEE 16 th International Symposium on Biomedical Imaging (ISBI 2019). IEEE, 2019: 953-956. [27] Lizzi F L, Astor M, Liu T, et al. Ultrasonic spectrum analysis for tissue assays and therapy evaluation [J]. International Journal of Imaging Systems and Technology, 1997, 8(1): 3-10. doi: 10.1002/(SICI)1098-1098(1997)8:1<3::AID-IMA2>3.0.CO;2-E [28] Linzer M, Norton S J. Ultrasonic tissue characterization [J]. Annual Review of Biophysics and Bioengineering, 1982, 11(1): 303-329. doi: 10.1146/annurev.bb.11.060182.001511 [29] Xu M, Wang L V. Photoacoustic imaging in biomedicine [J]. Review of Scientific Instruments, 2006, 77(4): 041101. doi: 10.1063/1.2195024 [30] Diem M, Chiriboga L, Yee H. Infrared spectroscopy of human cells and tissue. VIII. Strategies for analysis of infrared tissue mapping data and applications to liver tissue [J]. Biopolymers: Original Research on Biomolecules, 2000, 57(5): 282-290. [31] Xiao X, Ma L. Color transfer in correlated color space[C]//Proceedings of the 2006 ACM International Conference on Virtual Reality Continuum and Its Applications, 2006: 305-309. [32] Reinhard E, Adhikhmin M, Gooch B, et al. Color transfer between images [J]. IEEE Computer Graphics and Applications, 2001, 21(5): 34-41. [33] Welsh T, Ashikhmin M, Mueller K. Transferring color to greyscale images[C]//Proceedings of the 29 th Annual Conference on Computer Graphics and Interactive Techniques, 2002: 277-280. [34] Byra M, Galperin M, Ojeda-Fournier H, et al. Breast mass classification in sonography with transfer learning using a deep convolutional neural network and color conversion [J]. Medical Physics, 2019, 46(2): 746-755. doi: 10.1002/mp.13361 [35] Hwang Y, Lee J Y, So Kweon I, et al. Color transfer using probabilistic moving least squares[C]//Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, 2014: 3342-3349. [36] Jing Y, Yang Y, Feng Z, et al. Neural style transfer: A review [J]. IEEE Transactions on Visualization and Computer Graphics, 2019, 26(11): 3365-3385. [37] Wan S, Xia Y, Qi L, et al. Automated colorization of a grayscale image with seed points propagation [J]. IEEE Transactions on Multimedia, 2020, 22(7): 1756-1768. doi: 10.1109/TMM.2020.2976573 [38] Gatys L A, Ecker A S, Bethge M. A neural algorithm of artistic style [J]. arXiv preprint, 2015: 1508.06576. [39] Albawi S, Mohammed T A, Al-Zawi S. Understanding of a convolutional neural network[C]//2017 International Conference on Engineering and Technology (ICET). IEEE, 2017: 1-6. [40] Szegedy C, Liu W, Jia Y, et al. Going deeper with convolutions[C]//Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, 2015: 1-9. [41] Long J, Shelhamer E, Darrell T. Fully convolutional networks for semantic segmentation[C]//Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, 2015: 3431-3440. [42] Huang G, Liu Z, Van Der Maaten L, et al. Densely connected convolutional networks[C]//Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, 2017: 4700-4708. [43] Simonyan K, Zisserman A. Very deep convolutional networks for large-scale image recognition [J]. arXiv preprint, 2014: 1409.1556. [44] Krizhevsky A, Sutskever I, Hinton G E. Imagenet classification with deep convolutional neural networks [J]. Advances in Neural Information Processing Systems, 2012, 25: 1097-1105. [45] Li Y, Hao Z B, Lei H. Survey of convolutional neural network [J]. Journal of Computer Applications, 2016, 36(9): 2508-2515. [46] Pineda F J. Generalization of back-propagation to recurrent neural networks [J]. Physical Review Letters, 1987, 59(19): 2229. doi: 10.1103/PhysRevLett.59.2229 [47] Johnson J, Alahi A, Li Feifei. Perceptual losses for real-time style transfer and super-resolution[C]//European Conference on Computer Vision. Cham: Springer, 2016: 694-711. [48] Liu X C, Cheng M M, Lai Y K, et al. Depth-aware neural style transfer[C]//Proceedings of the Symposium on Non-Photorealistic Animation and Rendering, 2017: 1-10. [49] Goodfellow I, Pouget-Abadie J, Mirza M, et al. Generative adversarial nets[C]//Advances in Neural Information Processing Systems, 2014 : 2672-2680. [50] Liu H, Fu Z, Han J, et al. Single satellite imagery simultaneous super-resolution and colorization using multi-task deep neural networks [J]. Journal of Visual Communication and Image Representation, 2018, 53: 20-30. doi: 10.1016/j.jvcir.2018.02.016 [51] Izadyyazdanabadi M, Belykh E, Zhao X, et al. Fluorescence image histology pattern transformation using image style transfer [J]. Frontiers in Oncology, 2019, 9: 519. doi: 10.3389/fonc.2019.00519 [52] Rivenson Y, Wang H, Wei Z, et al. Virtual histological staining of unlabelled tissue-autofluorescence images via deep learning [J]. Nature Biomedical Engineering, 2019, 3(6): 466-477. doi: 10.1038/s41551-019-0362-y [53] Rivenson Y, de Haan K, Wallace W D, et al. Emerging advances to transform histopathology using virtual staining [J]. BME Frontiers, 2020, 2020: 9647163. doi: 10.34133/2020/9647163 [54] Nishar H, Chavanke N, Singhal N. Histopathological stain transfer using style transfer network with adversarial loss[C]//International Conference on Medical Image Computing and Computer-Assisted Intervention. Cham: Springer, 2020: 330-340. [55] Touvron H, Douze M, Cord M, et al. Powers of layers for image-to-image translation [J]. arXiv preprint, 2020: 2008.05763. [56] Zuo Z, Xu Q, Zhang H, et al. Multimodal image-to-image translation via mutual information estimation and maximization [J]. arXiv preprint, 2020: 2008.03529. [57] Isola P, Zhu J Y, Zhou T, et al. Image-to-image translation with conditional adversarial networks[C]//Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, 2017: 1125-1134. [58] Dai B, Fidler S, Urtasun R, et al. Towards diverse and natural image descriptions via a conditional gan[C]//Proceedings of the IEEE International Conference on Computer Vision, 2017: 2970-2979. [59] Zhu J Y, Park T, Isola P, et al. Unpaired image-to-image translation using cycle-consistent adversarial networks[C]//Proceedings of the IEEE International Conference on Computer Vision, 2017: 2223-2232. [60] Almahairi A, Rajeshwar S, Sordoni A, et al. Augmented cyclegan: Learning many-to-many mappings from unpaired data[C]//International Conference on Machine Learning. PMLR, 2018: 195-204. [61] Tschuchnig M E, Oostingh G J, Gadermayr M. Generative adversarial networks in digital pathology: A survey on trends and future potential [J]. Patterns, 2020, 1(6): 100089. doi: 10.1016/j.patter.2020.100089 [62] Petriceks A H, Olivas J C, Srivastava S. Trends in geriatrics graduate medical education programs and positions, 2001 to 2018 [J]. Gerontology and Geriatric Medicine, 2018, 4: 2333721418777659. doi: 10.1177/2333721418777659 [63] Shen D, Wu G, Suk H I. Deep learning in medical image analysis [J]. Annual Review of Biomedical Engineering, 2017, 19: 221-248. doi: 10.1146/annurev-bioeng-071516-044442 [64] Barcia J J. The Giemsa stain: Its history and applications [J]. International Journal of Surgical Pathology, 2007, 15(3): 292-296. doi: 10.1177/1066896907302239 [65] Daykin M E, Hussey R S. Staining and histopathological techniques [J]. An Advanced Treatise on Meloidogyne, 1985, 2: 39-48. [66] Pradhan P, Meyer T, Vieth M, et al. Computational tissue staining of non-linear multimodal imaging using supervised and unsupervised deep learning [J]. Biomedical Optics Express, 2021, 12(4): 2280-2298. doi: 10.1364/BOE.415962 [67] Barlow H B. Unsupervised learning [J]. Neural Computation, 1989, 1(3): 295-311. doi: 10.1162/neco.1989.1.3.295 [68] Teramoto A, Yamada A, Tsukamoto T, et al. Mutual stain conversion between Giemsa and Papanicolaou in cytological images using cycle generative adversarial network [J]. Heliyon, 2021, 7(2): e06331. doi: 10.1016/j.heliyon.2021.e06331 [69] Padma S, PV R, Kante R, et al. A comparative study of Staining characteristics of Leishman-Geimsa cocktail and Papanicolaou stain in Cervical Cytology [J]. Asian Pacific Journal of Health Sciences, 2018, 5: 233-236. doi: 10.21276/apjhs.2018.5.3.32 [70] Gollapudi B, Kamra O P. Applications of a simple Giemsa-staining method in the micronucleus test [J]. Mutat Res, 1979, 64(1): 45-46. doi: 10.1016/0165-1161(79)90135-3 [71] Woolman M, Tata A, Bluemke E, et al. An assessment of the utility of tissue smears in rapid cancer profiling with desorption electrospray ionization mass spectrometry (DESI-MS) [J]. Journal of The American Society for Mass Spectrometry, 2016, 28(1): 145-153. [72] Niazi M K K, Parwani A V, Gurcan M N. Digital pathology and artificial intelligence [J]. The Lancet Oncology, 2019, 20(5): e253-e261. doi: 10.1016/S1470-2045(19)30154-8 [73] Lo Y C, Chung I F, Guo S N, et al. Cycle-consistent GAN-based stain translation of renal pathology images with glomerulus detection application [J]. Applied Soft Computing, 2021, 98: 106822. doi: 10.1016/j.asoc.2020.106822 [74] Khojasteh M, Ward R, MacAulay C. Quantification of membrane IHC stains through multi-spectral imaging[C]//2012 9th IEEE International Symposium on Biomedical Imaging (ISBI). IEEE, 2012: 752-755. [75] Xu Z, Moro C F, Bozóky B, et al. GAN-based virtual re-staining: A promising solution for whole slide image analysis [J]. arXiv preprint, 2019: 1901.04059. [76] de Bel T, Bokhorst J M, van der Laak J, et al. Residual cyclegan for robust domain transformation of histopathological tissue slides [J]. Medical Image Analysis, 2021, 70: 102004. doi: 10.1016/j.media.2021.102004 [77] Bornstein M B. Reconstituted rat-tail collagen used as substrate for tissue cultures on coverslips in Maximow slides and roller tubes [J]. Laboratory Investigation, 1958, 7(2): 134-137. [78] Boonstra H, Oosterhuis J W, Oosterhuis A M, et al. Cervical tissue shrinkage by formaldehyde fixation, paraffin wax embedding, section cutting and mounting [J]. Virchows Archiv A, 1983, 402(2): 195-201. doi: 10.1007/BF00695061 [79] Jensen E C. Quantitative analysis of histological staining and fluorescence using ImageJ [J]. The Anatomical Record, 2013, 296(3): 378-381. doi: 10.1002/ar.22641 [80] Kang L, Li X, Zhang Y, et al. Deep learning enables ultraviolet photoacoustic microscopy based histological imaging with near real-time virtual staining [J]. Photoacoustics, 2021, 25: 100308. doi: 10.1016/j.pacs.2021.100308 [81] Baik J W, Kim H, Son M, et al. Intraoperative label-free photoacoustic histopathology of clinical specimens [J]. Laser & Photonics Reviews, 2021, 15(10): 2100124. [82] Morrison D, Harris-Birtill D, Caie P D. Generative deep learning in digital pathology workflows [J]. The American Journal of Pathology, 2021, 191(10): 1717-1723. doi: 10.1016/j.ajpath.2021.02.024 [83] Zhou N, Cai D, Han X, et al. Enhanced cycle-consistent generative adversarial network for color normalization of H&E stained images[C]//International Conference on Medical Image Computing and Computer-Assisted Intervention. Cham: Springer, 2019: 694-702. [84] Cho H, Lim S, Choi G, et al. Neural stain-style transfer learning using gan for histopathological images [J]. arXiv preprint, 2017: 1710.08543. [85] Li B, Keikhosravi A, Loeffler A G, et al. Single image super-resolution for whole slide image using convolutional neural networks and self-supervised color normalization [J]. Medical Image Analysis, 2021, 68: 101938. doi: 10.1016/j.media.2020.101938 [86] Chen X, Yu J, Cheng S, et al. An unsupervised style normalization method for cytopathology images [J]. Computational and Structural Biotechnology Journal, 2021, 19: 3852-3863. doi: 10.1016/j.csbj.2021.06.025 [87] Vahadane A, Peng T, Sethi A, et al. Structure-preserving color normalization and sparse stain separation for histological images [J]. IEEE Transactions on Medical Imaging, 2016, 35(8): 1962-1971. doi: 10.1109/TMI.2016.2529665 [88] Macenko M, Niethammer M, Marron J S, et al. A method for normalizing histology slides for quantitative analysis[C]//2009 IEEE International Symposium on Biomedical Imaging: From Nano to Macro. IEEE, 2009: 1107-1110. [89] Khan A M, Rajpoot N, Treanor D, et al. A nonlinear mapping approach to stain normalization in digital histopathology images using image-specific color deconvolution [J]. IEEE Transactions on Biomedical Engineering, 2014, 61(6): 1729-1738. doi: 10.1109/TBME.2014.2303294 [90] Gupta A, Duggal R, Gehlot S, et al. GCTI-SN: Geometry-inspired chemical and tissue invariant stain normalization of microscopic medical images [J]. Medical Image Analysis, 2020, 65: 101788. doi: 10.1016/j.media.2020.101788 [91] Zheng Y, Jiang Z, Zhang H, et al. Adaptive color deconvolution for histological WSI normalization [J]. Computer Methods and Programs in Biomedicine, 2019, 170: 107-120. doi: 10.1016/j.cmpb.2019.01.008 [92] Shaban M T, Baur C, Navab N, et al. Staingan: Stain style transfer for digital histological images[C]//2019 IEEE 16th International Symposium on Biomedical Imaging (ISBI 2019). IEEE, 2019: 953-956. [93] Tellez D, Litjens G, Bándi P, et al. Quantifying the effects of data augmentation and stain color normalization in convolutional neural networks for computational pathology [J]. Medical Image Analysis, 2019, 58: 101544. doi: 10.1016/j.media.2019.101544 [94] Bian Y, Jiang Y, Wang J, et al. Deep learning colorful ptychographic iterative engine lens-less diffraction microscopy [J]. Optics and Lasers in Engineering, 2022, 150: 106843. doi: 10.1016/j.optlaseng.2021.106843 [95] Shen H, Gao J. Deep learning virtual colorful lens-free on-chip microscopy [J]. Chinese Optics Letters, 2020, 18(12): 121705. doi: 10.3788/COL202018.121705 [96] Bian Y, Jiang Y, Huang Y, et al. Deep learning virtual colorization overcoming chromatic aberrations in singlet lens microscopy [J]. APL Photonics, 2021, 6(3): 031301. doi: 10.1063/5.0039206 -







 下载:
下载: